When it comes to down to it, the genomic variants we collect in a research and clinical setting are impossible to interpret without that important link of how genes are related to phenotypes.
Indisputably, the Johns Hopkins project to catalog all evidence related to inheritable Mendelian diseases is our best repository of this evidence.
Online Mendelian Inheritance in Man (OMIM) was founded in the ‘60s by the pioneering geneticist Dr. Vic McKusick at Johns Hopkins with the bold aim to be a full-text, comprehensive reference of all scientific evidence on the relationship between phenotype and genotype.
In our upcoming release of VarSeq, OMIM will be an add-on annotation source and will be deeply integrated with our soon-to-be-announced platform for generating clinical reports.
The OMIM Power Up
OMIM is tracking a fast moving target: all human knowledge on how genes function in humans.
With committed directorship of Dr Ada Hamosh (a name you will quickly be familiar with if looking at each page’s contributors list), OMIM is getting updated on a day to day basis with new literature, cross-references and hand-written descriptions and interpretations of genes, phenotypes and variants.
If you have noticed OMIM/MIM links in VarSeq or GenomeBrowse already, you are picking up on the prolific linkage other annotations sources provide to this canonical resource.
With the careful curation of the OMIM resource at the Phenotype, Gene and Variant level, the following descriptive and filterable data points will be joinable to your imported variants:
- Functional description of genes and phenotypes ready for physicians to orient themselves
- Lists of phenotypes linked to genes with supporting evidence and modes of inheritance of the phenotype (autosomal dominant, etc.)
- Paper references with relevant PubMed and direct URLs
- Links to clinical relevant genetic resources such as testing guidelines, ontology and gene test registries
- Descriptive interpretations for variants curated from published papers with family and disease context
For example, in this TruSight Cardio gene panel, I added a filter on the diseases linked to the gene and focused in on those shown to be linked to Cardiomyopathy.
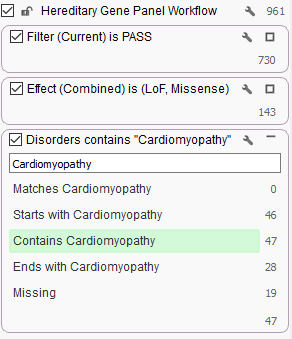
We will soon be posting about our plans to provide a flexible and powerful platform for authoring and generating clinical reports for lab test providers to use right within VarSeq.
With the full depth of the OMIM database linked in, we can provide very rich gene and phenotype clinical descriptions, variant interpretations and enumerated paper references for the variants selected to include in the report.




That’s a great move Gabe.
Back in action Nov 1 hoping there will be an update of VarSeq.
Dave
Any chance you’ll be adding OMIA for your non-human focused users?
Hi Matt,
Currently OMIA isn’t available in a user-friendly format, but if they put it into a more easily accessible format then we would certainly curate it and provide it to our users. So hopefully that is something on their to do list!
Will the platform for generating clinical reports be included in VarSeq?
Yes! Stay tuned for more announcements on this in the coming weeks.
Hi Gabe
When do you expect the new OMIM annotation to be available in VarSeq/GenomeBrowse?
Hi Martin,
OMIM is now available in VarSeq! You can read more about it here: https://www.goldenhelix.com/VarSeq/VarSeq_Reports.html