
Managing long‑read secondary analysis used to mean wrestling with Cromwell servers, custom compute nodes, and a tangle of WDL files. VSWarehouse 3 removes that burden. Drop your PacBio output into a watched folder, and our platform does the rest: spinning up compute, tracking every task, and handing you fully annotated results ready for clinical interpretation.
Built‑In Workflows, Ready to Launch
To provide this experience, we have brought the PacBio first-class workflows to the VSWarehouse platform.
These VSWarehouse workflows are built on PacBio’s own pipelines, containerized and version‑locked, so your lab gets identical results to PacBio’s white‑papers without any local engineering:
| Sample Type | Pipeline | Primary Outputs Captured |
| HiFi Whole Genome | HiFi‑human‑WGS‑WDL | Haplotagged BAM, coverage metrics, phased SNV/INDEL VCF, gVCF, structural‑variant VCF, Tandem‑Repeat (TRGT) VCF, methylation tracks, QC summary JSONs |
| PureTarget Carrier Panels | PureTarget Carrier Pipeline | pbmm2‑aligned BAM, panel‑aware TRGT VCF, Paraphase phased BAM/VCF, inversion calls, per‑sample QC summaries |
In VSWarehouse 3 you can simply download these workflows from our Workflow Registry. There are no prerequisite installs and no YAML editing. You select the input folder, the output folder, and hit Run.
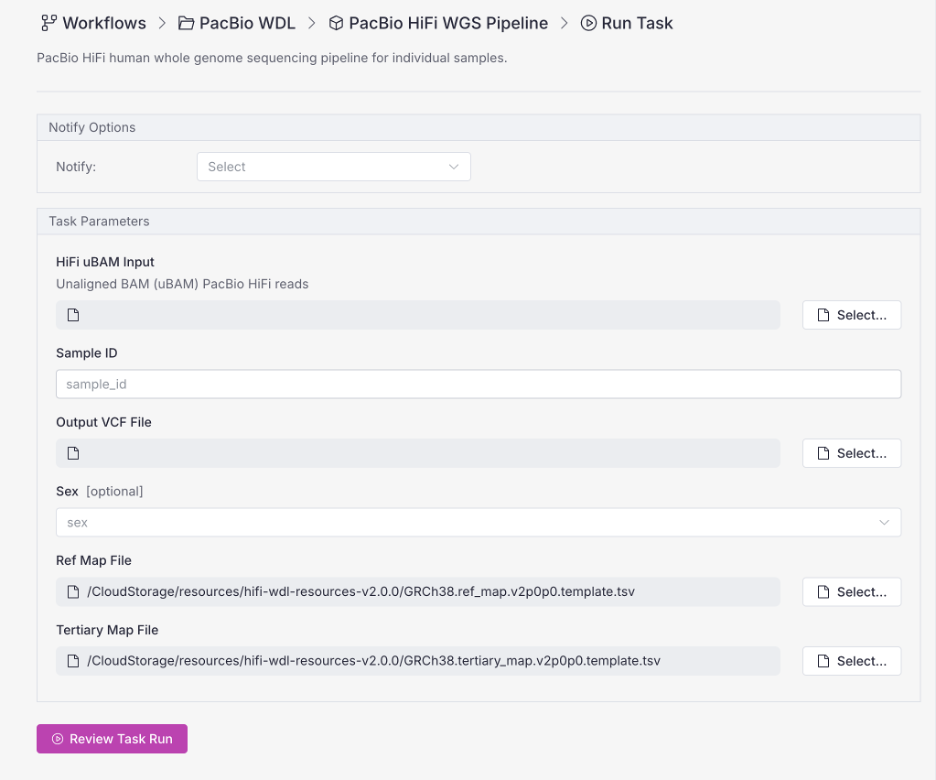
Seamless from Sequencer to Report
To get this experience, there are a couple of capabilities of VSWarehouse3 that are worth covering, and how they support the PacBio workflows:
- Auto‑Ingest: Define a watched file pattern and a parent path. New folders of uBAMs instantly trigger the correct pipeline.
- Resource‑Aware Scheduling: On-prem or transient cloud VMs, picked automatically according to the pipeline’s needs.
- Tertiary Analysis Integration: Successful secondary runs call VSPipeline with your VarSeq project template, creating a ready-to-interpret project and draft clinical reports.
- Notifications Your Way: Set up who gets notified when an automated pipeline completes so you know when to jump into the next step.
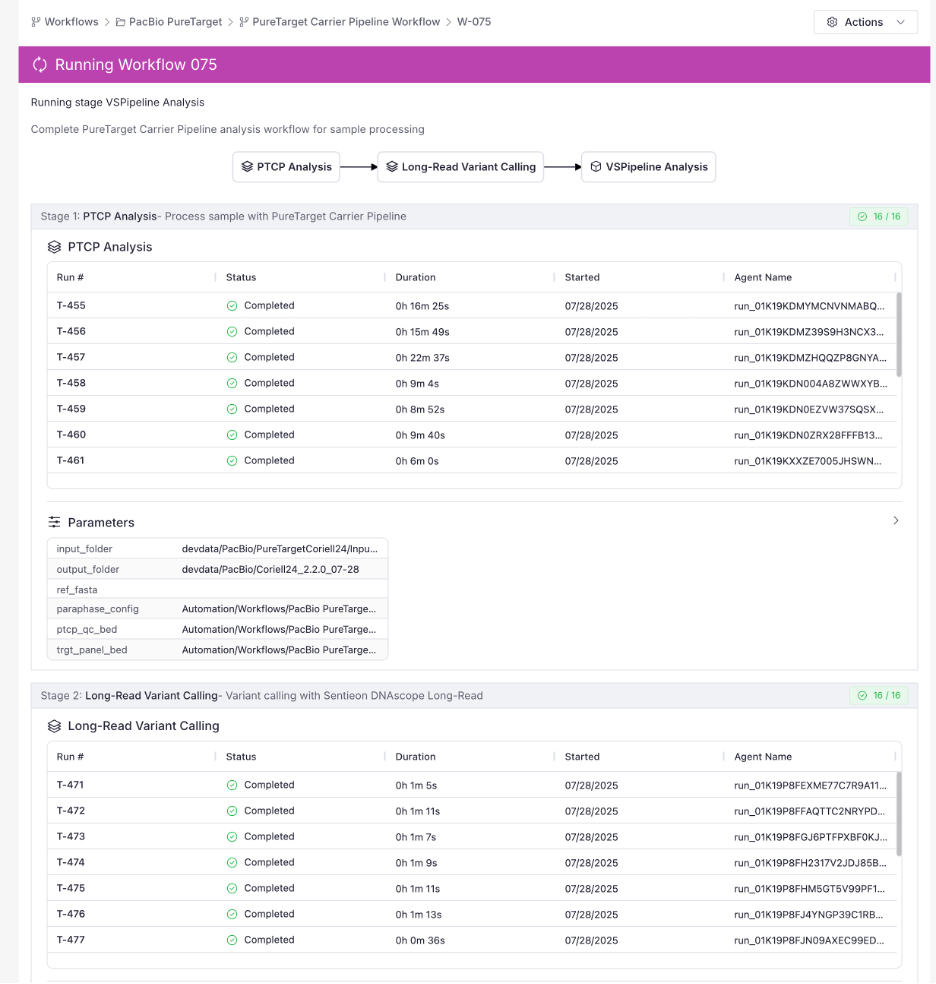
Transparent Monitoring
The Workflow and Task Run Monitor shows every stage, duration, and active task (see screenshot above). Need logs? Click the row, view stdout/stderr in‑browser in real time as the task is running. You can also download the full execution bundle, perfect for audits and troubleshooting.
Fits Your Infrastructure – Cloud or Local
Whether you wish to leverage cloud-native compute and storage scaling or the local infrastructure available to you, you can use the same VSWarehouse workflows with the workflow runner that looks and works the same.
Because it is based on Docker images and dynamic scaling, you can rely on reproducible results that efficiently utilize your resources. This is especially important as data sizes grow to whole genomes and sample counts increase.
If you are interested in evaluating VSWarehouse 3 with your PacBio data, reach out to us! We are happy to discuss your use case and tailor a test instance to your needs so you can see the pairing of automated secondary with the power of the VarSeq Suite for tertiary interpretation.





Leave a comment