In the rapidly evolving field of pharmacogenomics (PGx), laboratories often face a common challenge: balancing standardized guidelines with the need for specialized, lab-specific data. In the past, maintaining custom PGx annotations was difficult and time-consuming, with custom annotation tracks generated using bespoke curation scripts that merged internal data with VarSeq’s default annotations. In this blog post, we discuss how you can leverage custom PGx catalogs in VSWarehouse, to more easily create and manage specialized PGx annotations.
The Challenge of Custom PGx Annotations
VarSeq’s PGx Variant Detection and Recommendations algorithm is powered by six different annotation tracks that work together to support comprehensive PGx reporting. These tracks specify critical information including allele definitions, gene-drug relationships, and drug dosing recommendations.
While VarSeq ships with default annotation tracks that cover standard CPIC and FDA recommendations, many laboratories require specialized annotations. Whether it’s incorporating institution-specific genes, adding custom drugs, defining novel alleles, or implementing tailored recommendations, these customizations are often essential for meeting the unique needs of different organizations.
Until now, maintaining these customized annotations has been a complex and labor-intensive process. Labs needed to create new custom annotation files using specialized curation scripts that merged their proprietary data with VarSeq’s default annotation tracks. This approach required technical expertise and was time-consuming to maintain.
PGx Catalogs in VSWarehouse
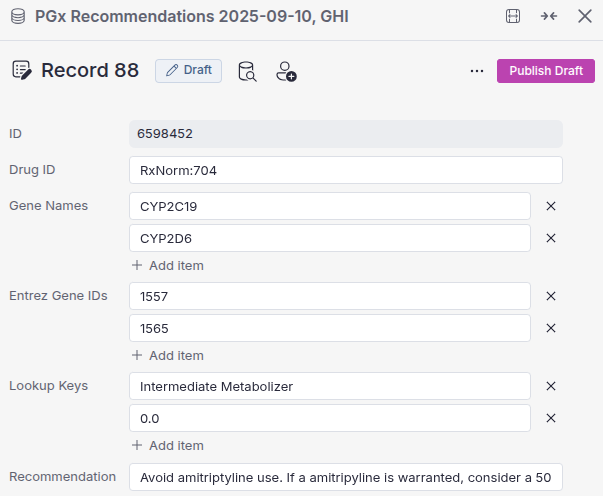
VSWarehouse users now have a streamlined alternative. By leveraging custom PGx Catalogs, you can create and manage specialized PGx annotations directly within VSWarehouse. These catalogs are stored centrally on the warehouse and can be seamlessly utilized by VarSeq to perform PGx annotation, just like the default annotation tracks that ship with the software.
Initially, PGx catalogs are created by importing data directly from VarSeq’s default CPIC and FDA annotation tracks. These catalogs can then be managed and updated directly through VSWarehouse’s intuitive catalog management interface, eliminating the need for custom scripting entirely.
This means your lab can easily add or modify genes, drugs, alleles, and recommendations as clinical guidelines evolve or institutional requirements change.
Powerful Search Capabilities
The catalog management system doesn’t just make it easy to maintain your annotations, it also provides robust search capabilities to help you quickly find and review specific information within your catalogs.
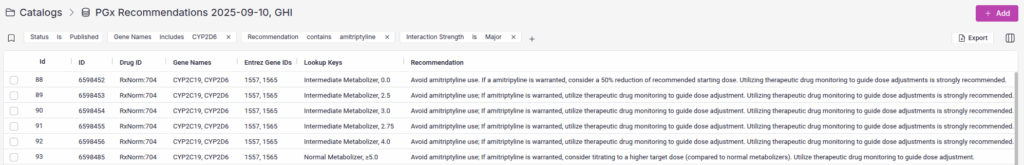
For example, you can query all recommendations for a specific drug that involve major interactions with a particular gene. In the screenshot below, we’re searching for all amitriptyline recommendations that have major drug interactions with CYP2D6, instantly surfacing the relevant clinical guidance.
This search functionality makes it simple to validate your annotations, audit your recommendations, and quickly locate specific diplotypes or gene-drug pairs during catalog development or review.
Getting Started with Custom PGx Catalogs
If you’re interested in learning more, or would like help getting started with custom PGx catalogs, please reach out to us. Our team can work with you to get your custom catalogs set up and integrated into your existing VarSeq workflows.