Explore the importance of negative findings in genomic medicine through the lens of VarSeq’s VSClinical AMP.
When analyzing a somatic sample in VarSeq, users have the option to report on several types of biomarkers with VSClinical AMP. In addition to your usual mutational biomarkers such as small variants, your copy number variants, and structural variants, we support the analysis of genomic signatures like microsatellite instability, tumor mutation burden, and a biomarker type that is not a mutation at all but what we call a negative finding. This blog will cover reporting negative findings biomarkers.
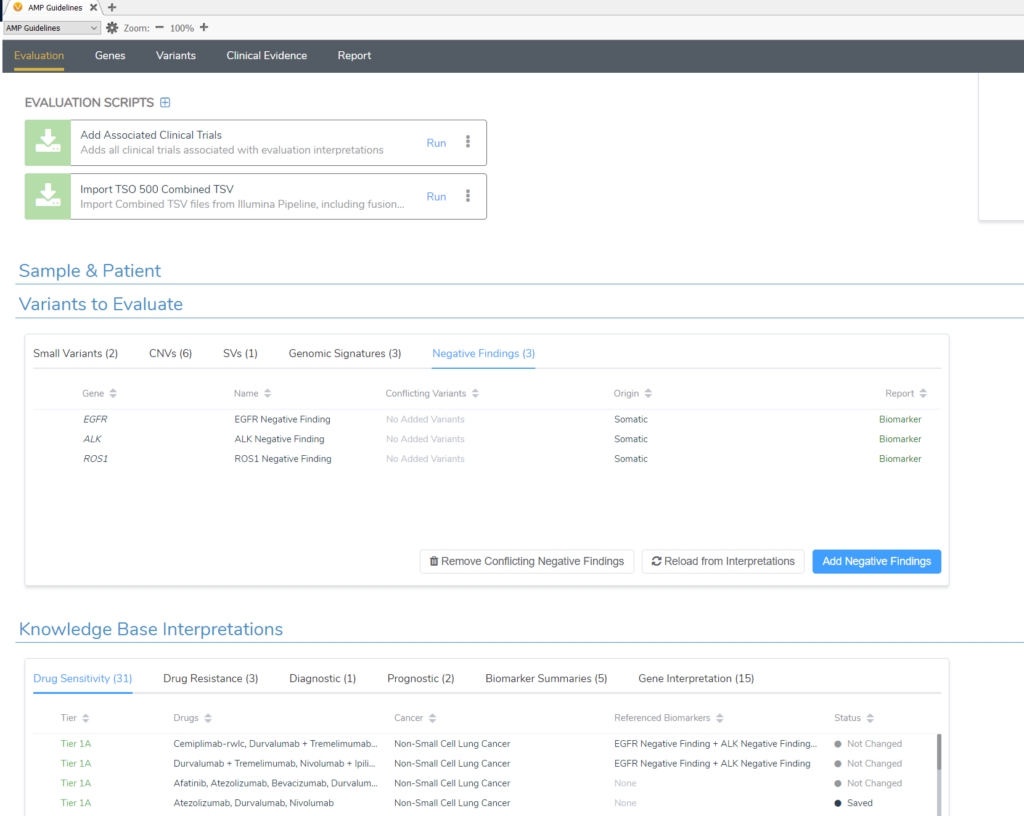
The term negative findings may sound unusual at first, but let us offer some clarity. Negative findings, simply put, refer to the state where a patient has wild-type alleles of the gene in question when that wild-type status is clinically significant for the selected cancer type. They are special biomarker types that are added in VSClinical automatically from our Golden Helix CancerKB database and are not variants filtered from your VCF. For example, if you select Lung Adenocarcinoma for the tumor type for evaluation, Golden Helix CancerKB will automatically add the negative finding biomarker for ALK and others like EGFR in the Variants to Evaluate tab (Figure 1). This is because, based on the clinical evidence for lung cancer patients, having wild-type ALK or EGFR is of specific clinical diagnostic and prognostic significance. In the figure below, you can see we automatically list the drugs that a patient with negative findings (wild type) in EGFR and ALK could be sensitive to. If you do have a variant to report in one of these genes, then the negative finding record for that gene should be removed.
Below is an example of the Golden Helix CancerKB curated drug sensitivity interpretation for an EGFR and ALK-negative finding. This shows that the drug combination nivolumab+ipilimumab along with chemotherapy is FDA-approved for NSCLC patients with wild-type EGFR and ALK, having been shown to improve the overall survival of similar patients regardless of their PD-L1 status in at least two clinical studies. On the other hand, a patient having a variant in EGFR or ALK would not be eligible for this drug combination.
VSClinical AMP, with our Golden Helix CancerKB knowledgebase, provides a very comprehensive somatic variant interpretation. We are not limited to only mutational biomarkers, and so give users the ability to find and report much relevant information on drugs that may be highly beneficial to a patient in need of treatment. For any questions or comments on this topic, please reach out to [email protected].