
Breakends (BNDs) represent the precise genomic coordinates where DNA is rearranged in structural variants such as translocations, inversions, and complex rearrangements. Analyzing breakends in NGS analysis is important because they capture the exact “breakpoints” of structural alterations, enabling accurate identification of gene fusions, large deletions, or other events that can disrupt or activate genes of clinical relevance. Systematically tracking how often breakends occur across samples further helps distinguish sequencing artifacts from true biological signals, revealing recurrent hotspots of genomic instability or clinically significant rearrangements.
Breakend Cataloging in VarSeq
VarSeq first introduced support for importing and analyzing breakends in version 2.3.0, but users could only store breakend cases in VSClinical catalogs at the gene or region level. This approach was helpful for annotation and clinical evaluation of recurring gene fusions but lacked coordinate-level precision, making it harder to track specific recurring events.
With the release of VarSeq 3.0, coordinate-aware breakend cataloging is now possible. This advancement allows users to store and query breakends at the exact genomic positions where the structural junction occurs, creating a more powerful way to identify recurrent events across projects.
How the New Catalogs Work
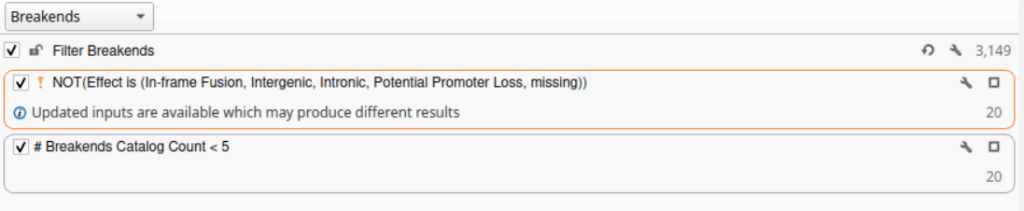
The new breakend catalogs are unique in that each record must capture two genomic coordinates, representing both sides of a structural junction. As such, the new catalogs include the following key fields: Chromosome (Chr), position (Start/Stop), retention direction (Retained), and inserted sequence (Insertion). Together, these fields preserve complete breakpoint context while maintaining compatibility with standard genomic position formats. Additional custom fields can also be added to support annotation and interpretation workflows. By simplifying the structure while retaining key information, these catalogs provide an efficient framework for tracking clinically relevant breakends and identifying potential artifacts.
Conclusion
With coordinate-aware breakend cataloging, VarSeq 3.0 gives researchers and clinicians a more precise and scalable way to interpret structural variants. By storing both sides of each junction and tracking their recurrence across samples, this functionality enables users to better separate noise from meaningful events. To learn more about the new breakend catalogs or evaluate VarSeq 3.0 please reach out to us at [email protected].



