
Here at Golden Helix, we continue to develop top-quality bioinformatic software to support high-throughput clinical next-gen sequencing pipelines, including long-read whole-genome-sequencing (WGS) workflows. WGS long-read technologies like PacBio HiFi and Oxford Nanopore offer improved variant calling for SNVs, Indels, structural variants, tandem repeats and repeat expansions, and epigenetic modifications. Each of these genetic variations that are detected using long-read WGS technology have important clinical applications including rare disease diagnostics, prenatal testing, and even oncology accentuating the need for high-quality data analysis tools and databases to aid in capturing the clinical impact of these variants.
VarSeq integrates long-read sequencing data from providers like PacBio and Oxford Nanopore and now we have expanded the resources available to assess the clinical impact of these variants by making the Consortium of Long Read Sequencing (CoLoRS) available within VarSeq for both GRCh37 and GRCh38-based workflows .
For those who are just starting to dabble in long-read analysis, the CoLoRS Database is a publicly accessible variant-frequency database derived from long‑read whole-genome sequencing (WGS), particularly PacBio HiFi reads. The data within CoLoRSDB is aggregated from nearly 1,400 human genomes, both rare-disease cohorts and population controls. It complements databases like gnomAD, which is built predominantly on short-read data, but captures variant types that short reads often miss such as structural variants (SVs), tandem repeats, small insertions/deletions (indels), and SNVs.
The figure below shows the advantage of including the CoLoRS database in your VarSeq long-read workflow. Plotted from top to bottom is RefSeq Genes, gnomAD variant frequencies, sample variants, CoLoRS variant frequencies, CoLoRS structural variant frequencies, then sample CNVs. The CoLoRSDB contains frequencies for a number of variants upstream of the ACSM2B gene including the C/A variant at Chr16: 20,530,570 – 20,530,570 in addition to a copy number deletion called for this sample that has frequency information from CoLoRSDB but are not captured within gnomAD. These variants were detected by long-read sequencing platforms and though little may be known about these variants as long-read sequencing continues to become widely used in clinical workflows, CoLoRSDB is providing insight to how common these newly detected variants are which is essential for assessing if these variations may be key drivers in rare diseases.
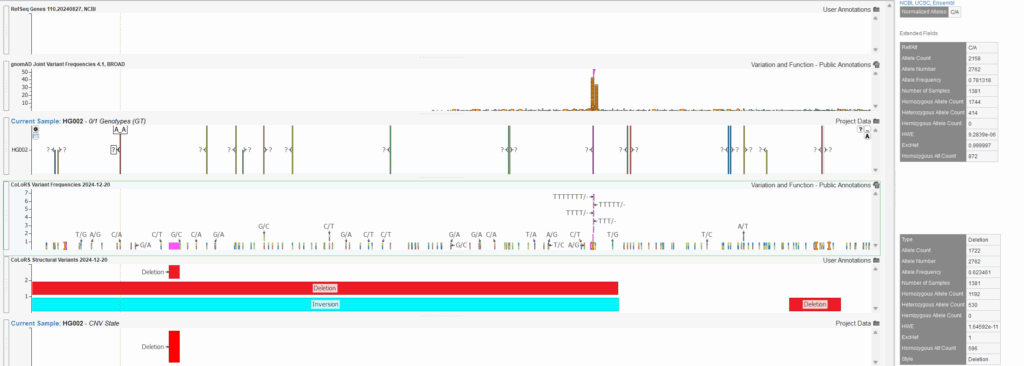
As whole-genome-sequencing becomes more routine in clinical workflows, databases like CoLoRSDB are essential to start assessing the clinical impact of these variants.
For more information or guidance on applying CoLoRSDB please do not hesitate to reach out to our team!



