Thank you to everyone who joined our recent webcast, “VSWarehouse for Genome Centers: Scalable, Secure Whole-Genome Infrastructure for Modern Sequencing Programs,” presented by Gabe Rudy on December 10, 2025. We appreciate the strong attendance and the excellent questions regarding the operational and security needs of modern genome centers.
For those who missed it or need a recap, the session focused on how VSWarehouse 3 and VarSeq 3 provide a single-tenant, highly secure platform capable of handling everything from individual clinical cases to population-scale cohorts. Whether deployed on-premises or in a private cloud, the platform addresses the challenge of scaling compute without compromising data sovereignty.
Meeting the Requirements of Modern Genome Centers
The webcast began by framing the core challenge: scaling from hundreds to tens of thousands of genomes requires consideration of resources, as well as data sovereignty and governance. VSWarehouse 3 is architected to meet these needs through flexible deployment models.
- Hybrid & Air-Gapped Deployment: We discussed the paradox of needing to scale while maintaining strict data control. VSWarehouse supports deployment on local high-performance computing (HPC) clusters for horizontal scaling, as well as on private clouds (AWS/Azure). Crucially, the platform fully supports air-gapped and highly restricted network environments, such as the solution currently deployed across the entire country of Denmark for its national genome center.
- Enterprise Security and Governance: To support multi-team environments, we showcased the “Workspaces” architecture. This allows a single instance to isolate data and users, separating cancer research from clinical rare disease, for example, while integrating with Active Directory/LDAP for seamless identity management and providing comprehensive audit trails.
- Population-Scale Knowledge: A major theme of the presentation was the transition from relying solely on public databases to building internal knowledge bases. While databases like gnomAD are essential, they often fail to capture local population-specificities or sequencing artifacts unique to a center’s library prep.
The webinar demonstrated a workflow processing over 3,100 whole genomes to build a local population allele frequency track. The demo highlighted the system’s ability to handle incremental updates: a new batch of 100 samples was processed and merged into the main cohort in just a few hours. This turns cohort management from a massive annual project into a routine background task, significantly reducing Variants of Unknown Significance (VUS) in downstream analysis.
Demo 1: Comprehensive Whole Genome Trio Analysis at Scale
The first deep-dive demonstration showcased a “FASTQ to Report” workflow for a Germline Trio running on an on-prem cluster. This workflow utilized Sentieon for highly efficient secondary analysis, including the calling of SNVs, CNVs, Structural Variants (SVs), and Tandem Repeats. The workflow then created per-trio VarSeq projects that integrated each of these specialized variant types into the analysis.
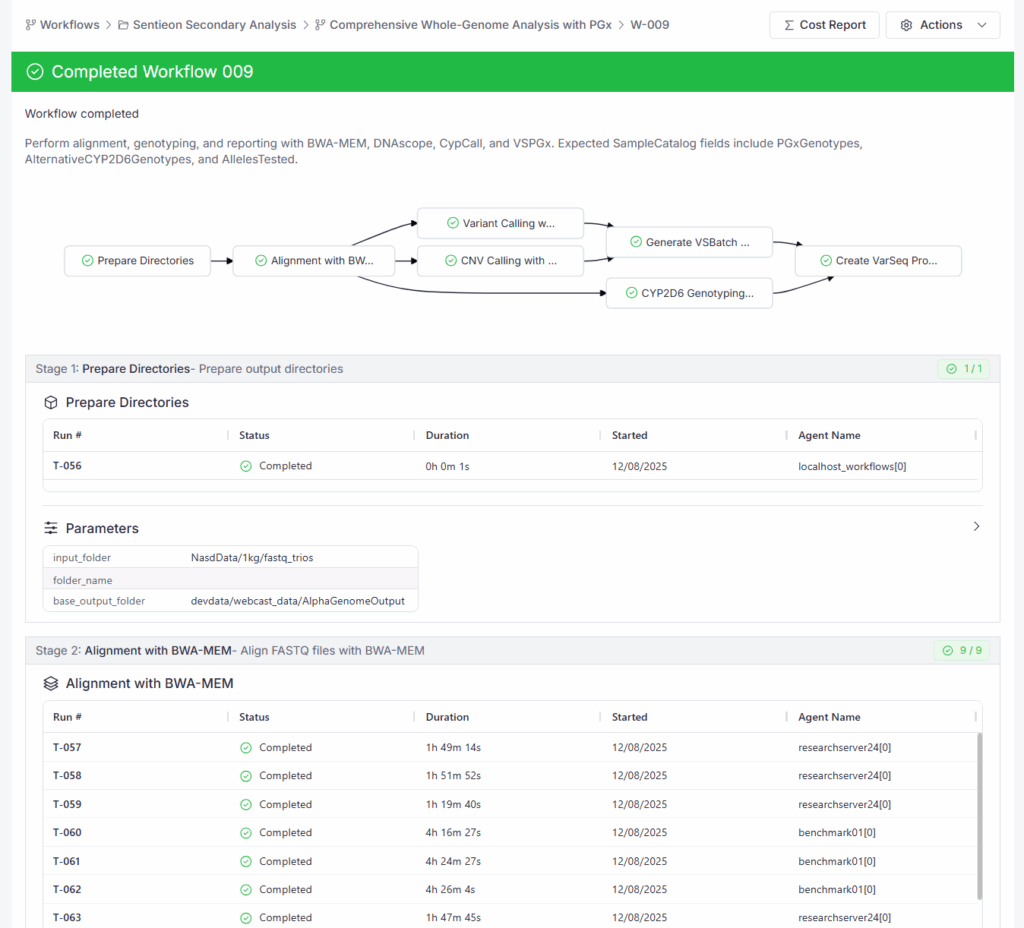
The Comprehensive Whole Genome Workflow calls small variants, CNVs, Structural Variants, and Tandem Repeats while automating the tertiary analysis with VarSeq
The power of this workflow lies in the integration of data types with the internal knowledge base and allele frequency cohorts. We demonstrated how the annotation and filtering reduced a list of 10 million raw variants to a manageable set of candidates by applying the ACMG auto-classifier, inheritance models, and internal frequency filters. Furthermore, the demo highlighted how Pharmacogenomics (PGx) data is automatically extracted, combining short-read calls and specialized genotypers (e.g., CYP2D6) to generate a comprehensive PGx report alongside the clinical findings.
Demo 2: Cloud-Native Tumor-Normal Workflows
Switching to an Azure-hosted environment, the demo illustrated a Somatic Whole Genome analysis. This highlighted the platform’s “Cloud-Native” capabilities:
- Dynamic Scaling: The system spun up 32-core virtual machines solely for the duration of the alignment and calling steps, shutting them down immediately upon completion.
- Cost Transparency: We viewed a granular cost report, revealing a compute cost of roughly $4.00 per whole genome sample pair.
- Advanced Visualization: We demonstrated the Stack-by-Base view in the GenomeBrowse interface. This feature moves reads with alternate alleles to the top of the pileup, making it effortless to inspect low-VAF somatic mutations and visualize phasing relationships.
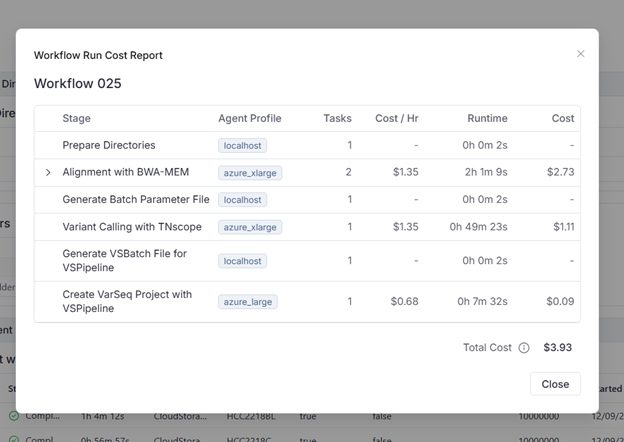
The whole genome tumor-normal workflow for a single WGS sample cost report
Somatic Variant Catalogs and Building on Golden Helix CancerKB
Finally, we covered how to automate the retention of institutional knowledge using Saved Exports. The demo showed how a completed somatic project can automatically push relevant data, such as tumor type, gene, and VAF, back into a central Somatic Assessment Catalog. This ensures that every future analysis benefits from the interpretations and observations of previous cases.
Our demo concluded with a tour through VSClinical AMP, where the candidate somatic variants were annotated with Golden Helix CancerKB and the institution’s cancer interpretations to report on drug recommendations, prognostic, and diagnostic evidence.
Looking Ahead
VSWarehouse and VarSeq 3 provide the secure, scalable foundation necessary for genome centers to operate with confidence. Whether you are running on-premises to meet strict regulatory needs or leveraging the elasticity of the cloud, the platform is designed to grow with your sequencing volume while building a permanent asset of genomic knowledge.
If you are considering scaling up your operations or consolidating multiple labs onto a unified platform, don’t hesitate to get in touch with our team to discuss a pilot deployment or an architecture review.



