
Effective use of data drives efficient, reliable analysis in next-generation sequencing. Users of the VarSeq suite are well-acquainted with the tools that VarSeq supplies to empower users, at the center of which lies a curated collection of clinical annotations that users can leverage in their own customized workflows. In the decades that we’ve improved and expanded the VarSeq suite since its release, we’ve also tracked the moving target of clinical knowledge. In 2026, users can explore 385 individual annotation tracks, consisting of 13.1 billion records — 22.5% of which weren’t in our database 5 years ago. More isn’t always better though; that’s why we employ a comprehensive curation process each month to ensure that what we’re shipping is consistent and applicable, and that new additions can be implemented into user workflows confidently, while giving users the autonomy to decide when and which tracks to update. Let’s break down some of the most poignant tracks in our library, and explore how they’ve improved not just in quantity but in quality over time, with more actionable records at our users’ fingertips.
What we curate
At a glance, our data source library, consisting of both open-source and proprietary tracks, consists of a growing list of consortia, registries, and peer-reviewed sources, with clinical mainstays like ClinVar, ClinGen, and CIViC, industry-standard gene tracks like OMIM, and specialized tracks like BRCA Exchange. In addition, we provide our own in-house, expert-curated Cancer Knowledge Base, which is the subject of previous and future detailed blog posts.
Different tracks come from different places and serve different purposes, and thus we see appropriate growth curves for individual tracks (Fig. 1). While knowledge bases like ClinVar represent the near-exponential growth of amalgamated industry expertise, other tracks like OMIM and ClinGen reflect growth not necessarily in the number of records, but in the quality of those records as information and actionability is augmented over time (Fig. 5).
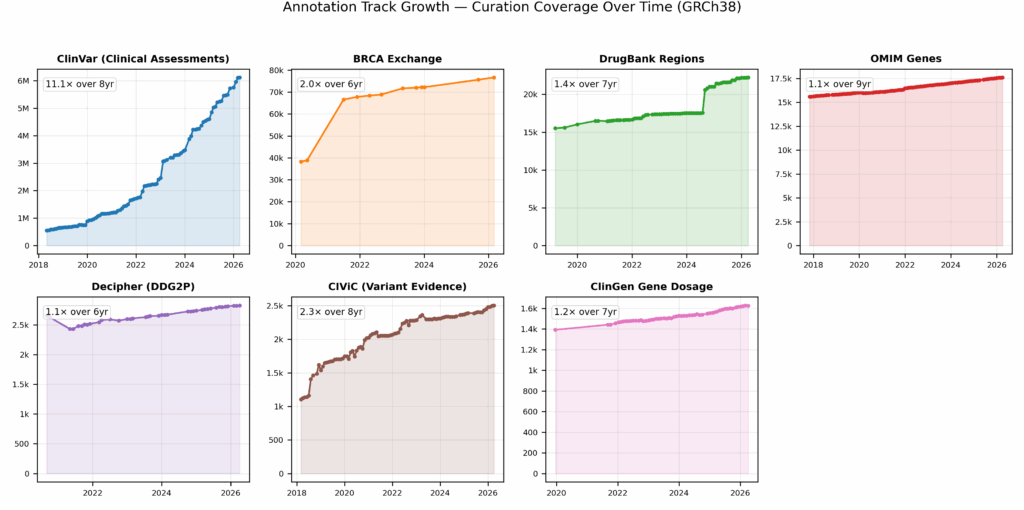
Small or large, we stay abreast of all changes made to tracks during our curation process. Smaller tracks may have fewer records, but often the individual records may prove more salient to our users investigating relevant issues (Fig. 2).
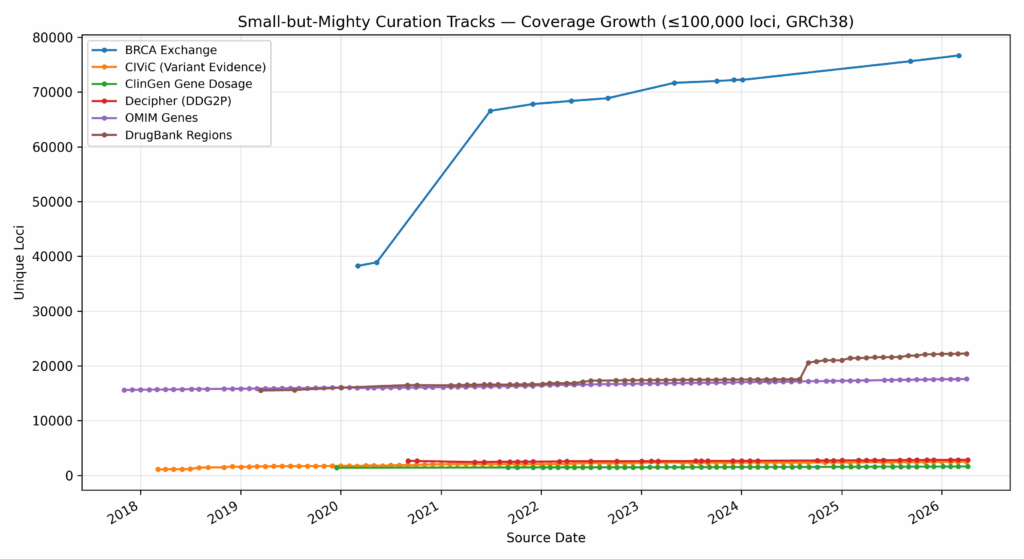
A window into clinical knowledge
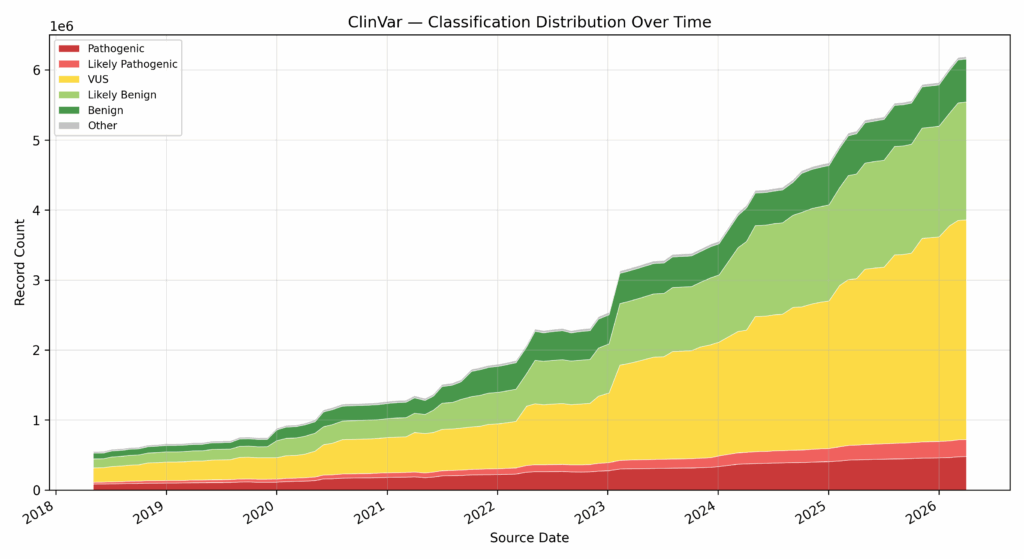
As we mentioned above, quantity doesn’t necessarily imply quality. The actionability of new records is the indicator of value. In the case of ClinVar, we can see that, while the total number of records has increased substantially, the number of benign, likely benign, pathogenic, and likely pathogenic variants have kept abreast of VUS submissions (Fig. 3). Each new submission adds evidence to user workflows that can be used to minimize the number of false positives on the benign side, and ensure capture of true positives in the pathogenic case. In the case of CIViC, we see a similar story: over time, the number of Level A — Validated and Level B — Clinical increases at pace with the overall number of records, even outpacing lower-impact submissions over the past few years (Fig. 4).
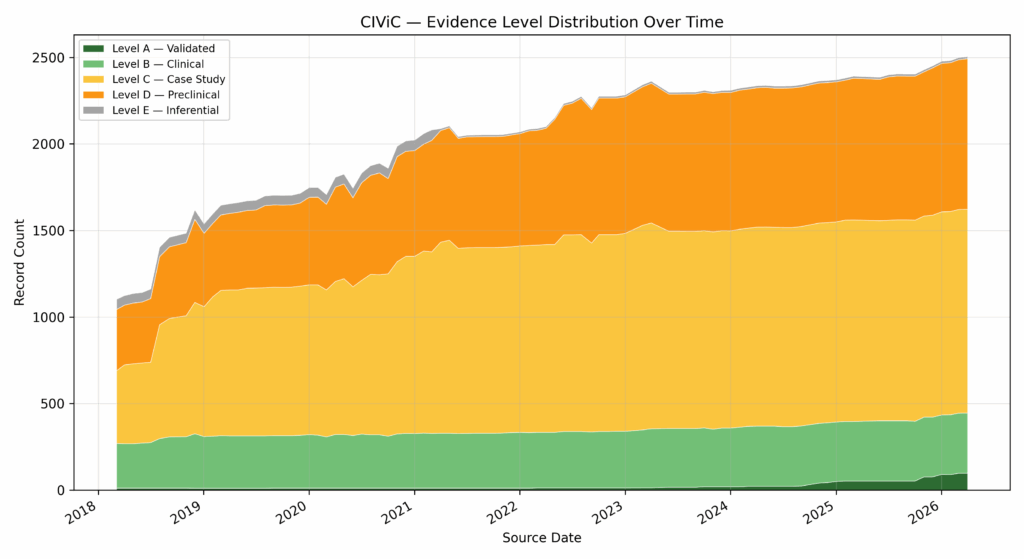
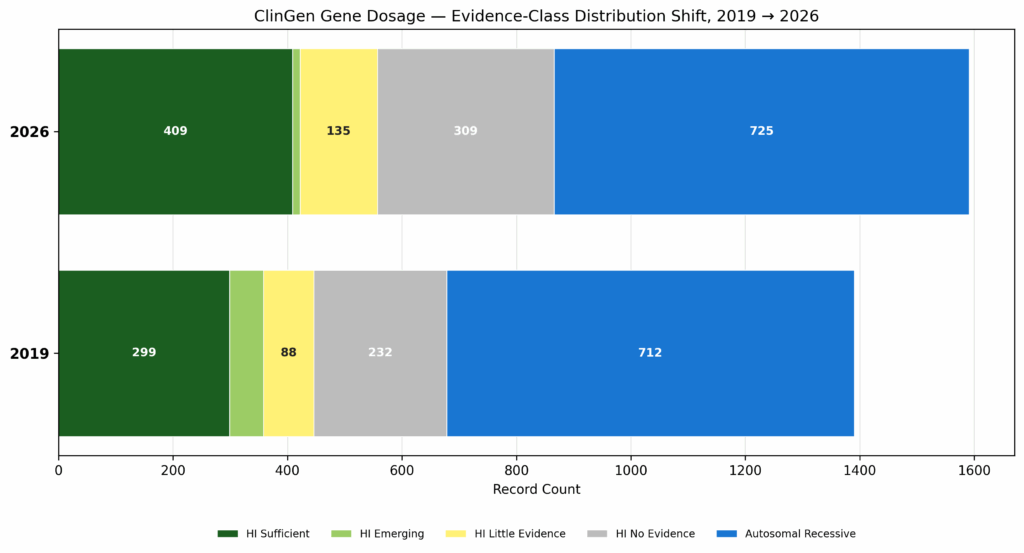
ClinGen Gene Dosage tells a similar story on the haploinsufficiency front. As the number of records increase, we see a higher ration of actionable haploinsufficiency and autosomal recessive submissions, decreasing the relative grey area of insufficient evidence (Fig. 5).
Bringing it all together
Naturally, we expect all of these results: legions of hard-working clinicians and researchers around the globe contribute to these tracks, so it’s no surprise that their utility increases over time. With the VarSeq suite, paired with our comprehensive curation efforts, however, users can access all of this new data automatically and seamlessly, without relying on their own curation process or manually querying records. Instead, existing workflows are updated when the user chooses to incorporate new track versions.
Questions, comments, or commendations for this post? Or is your workflow in need of an upgrade that allows you to automatically incorporate the vast amount of clinical knowledge available in our 300+ curated annotation tracks? Reach out to our team, we’re looking forward to hearing from you!