DNA methylation plays a critical role in gene regulation and is increasingly recognized as a clinically meaningful biomarker in oncology. With the rise of long-read sequencing, we now have the ability to detect base modifications alongside small variants and structural variations. In this post, we’ll walk through how VSWarehouse and VarSeq support methylation analysis, from raw PacBio reads all the way to clinical reporting.
A Somatic Workflow Built for Long Reads
VSWarehouse provides a PacBio WGS Somatic Workflow, available for download from our Workflow Repository. This automated pipeline handles everything from alignment to variant calling, phasing, structural variant detection, and methylation analysis, all from a single sequencing run.
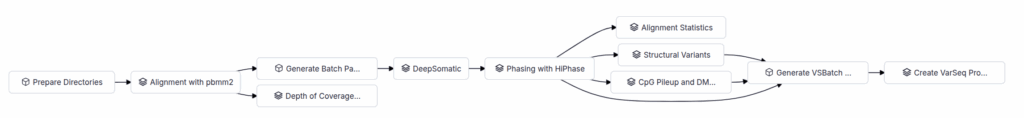
At the heart of our methylation support is the CpG Pileup and DMR Analysis stage, which performs two key tasks: generating per-CpG methylation scores across the genome, and identifying differentially methylated regions (DMRs) between tumor and matched normal samples.
Identifying Differentially Methylated Regions
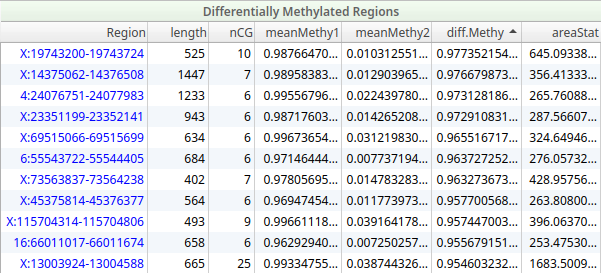
The DMR output captures the genomic coordinates of each region along with the number of CpG sites covered, mean methylation levels in the tumor and normal, the difference between them, and a statistical area score reflecting the strength of the signal. The mean methylation differences between tumor and normal tissue can point to biologically and potentially clinically significant epigenetic alterations.
DMRs can be imported directly into VarSeq, where they appear as a filterable table allowing users to sort and filter on methylation difference, region length, statistical significance, and genomic location, making it straightforward to prioritize the most meaningful epigenetic events.
Visualizing Methylation Data
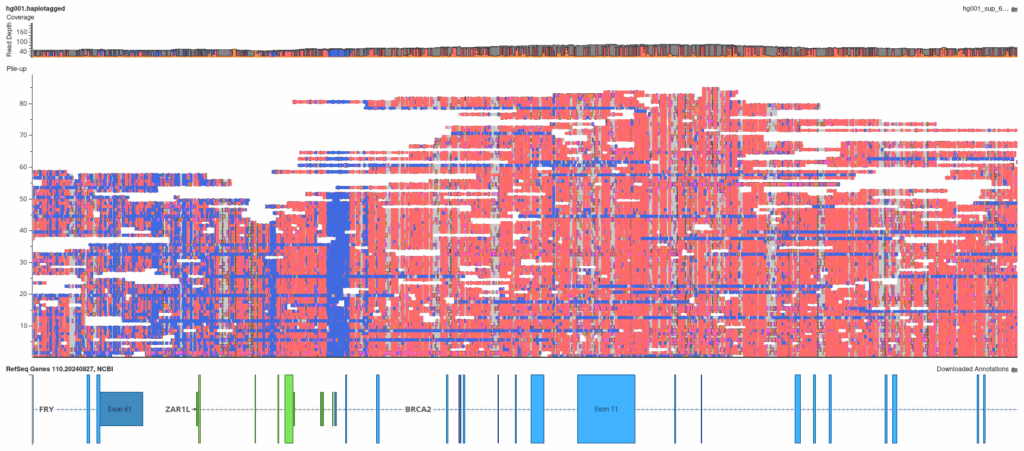
VarSeq’s integrated genome browser can render base modification data stored directly in long-read BAM files (one of the unique advantages of PacBio HiFi sequencing over short-read technologies). The coloring scheme is designed for immediate visual clarity: bases with a high probability of methylation are colored red, while bases with a low methylation probability are colored blue. This intuitive display makes it easy to spot regions of aberrant hypermethylation at single-base resolution.
Reporting Methylation Biomarkers in VSClinical
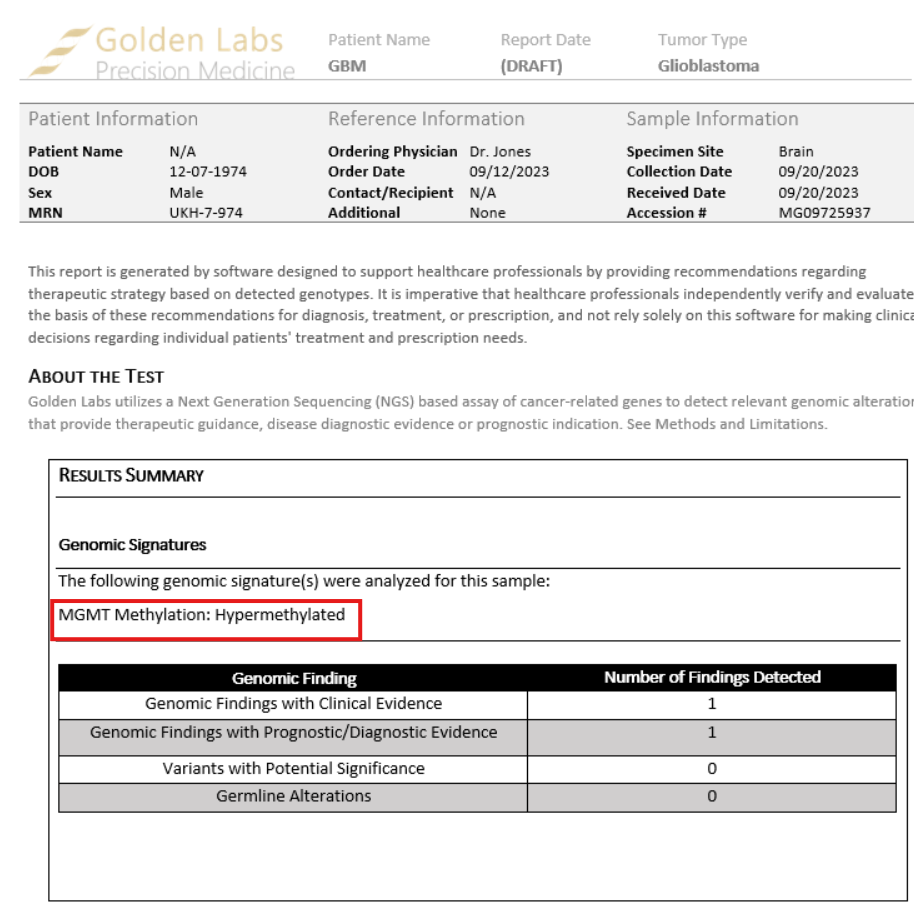
While our variant interpretation and reporting platform, VSClinical, is primarily oriented toward sequence variants, copy number variants, and structural variations, we recognize that epigenetic biomarkers are increasingly important in clinical oncology. A well-established example is MGMT promoter methylation in glioblastoma, which is both a prognostic marker and a predictor of response to temozolomide chemotherapy.
To support these use cases, VSClinical allows users to create and report user-defined interpretations for methylated DNA biomarkers, enabling labs to document and communicate clinically relevant methylation findings within their standard reporting workflow.
Conclusion
Long-read sequencing technology provides the unique ability to analyze both genetic variation and epigenetic modifications using the same dataset. Through our PacBio WGS Somatic Workflow, we are able to deliver a truly integrated view of the cancer genome, including sequence variants, structural variants, phasing, and epigenetics, all from a single sequencing run. If you’re interested in learning more about how VarSeq and VSWarehouse can support your long-read sequencing workflows, please reach out to our team using the contact link below.