
As long-read sequencing solidifies its role in clinical and translational research, labs are increasingly seeking streamlined methods to transition from raw data to actionable results. With the release of VSWarehouse 3.0, Golden Helix now supports a fully integrated implementation of PacBio’s somatic workflow, providing users with a powerful, centralized environment for managing large datasets, running best-practice pipelines, and automating tertiary interpretation, all without introducing procedural complexity.
A Unified Workspace for Long-Read Somatic Analysis
With VSWarehouse 3.0, teams can seamlessly run PacBio’s somatic pipeline directly within their workspace. As seen in Figures 1 & 2, this brings long-read alignment, DeepSomatic variant calling, and downstream annotation into a unified framework designed for reproducibility, version control, and collaboration. The result: labs can operationalize long-read somatic analysis the same way they already manage germline and short-read workflows with consistency, automation, and enterprise-grade audit trails.
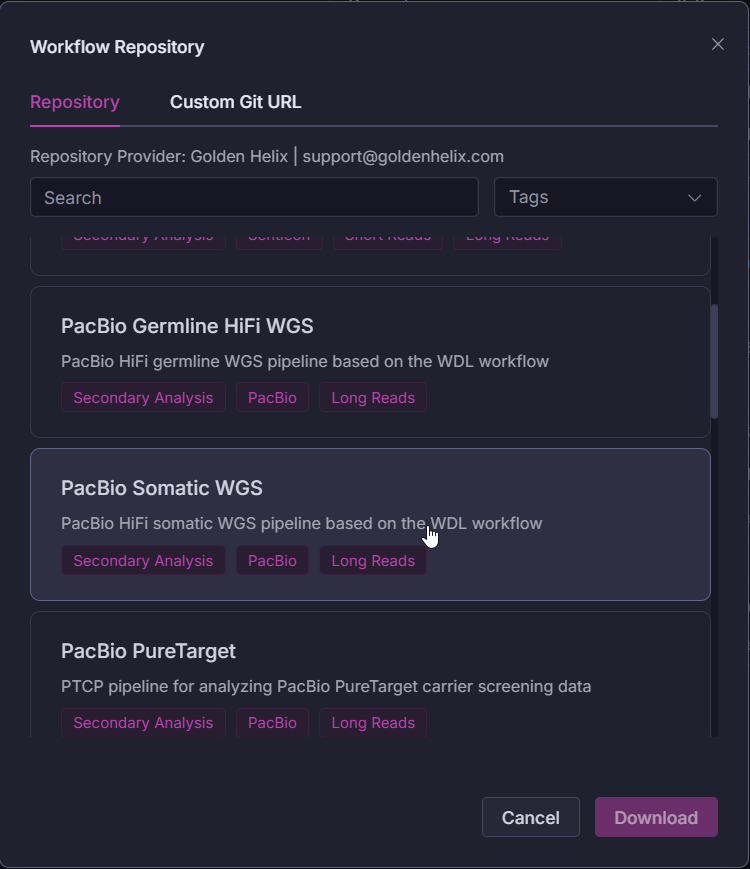
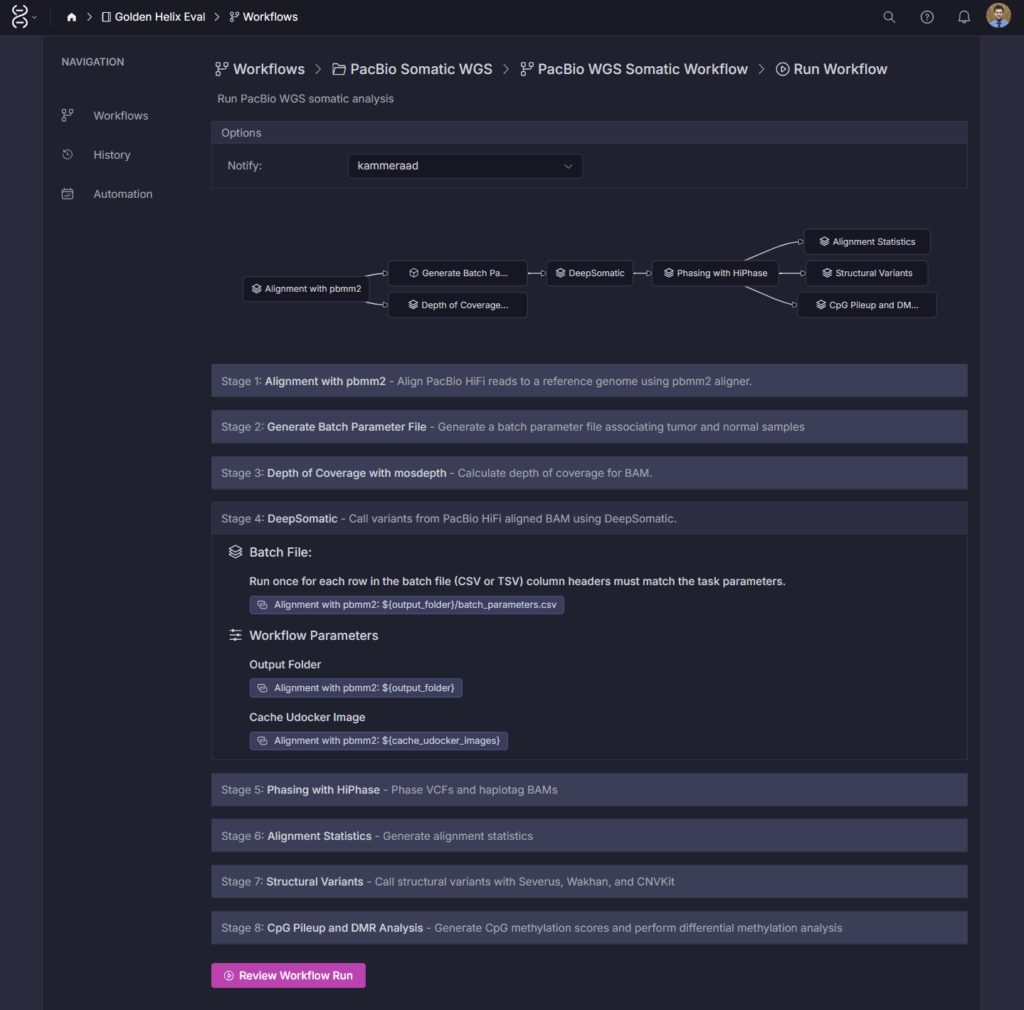
Streamlined, Automated Tertiary Interpretation
Once variants are called, the workflow naturally extends into VarSeq’s powerful tertiary analysis stack as seen in Figure 3. Users can apply templated filters, leverage curated annotation sources, and automatically generate clinical or research-grade reports. VSWarehouse ensures that all pipeline outputs, QC metrics, and interpretations are stored centrally, making data traceable and sharable across teams.
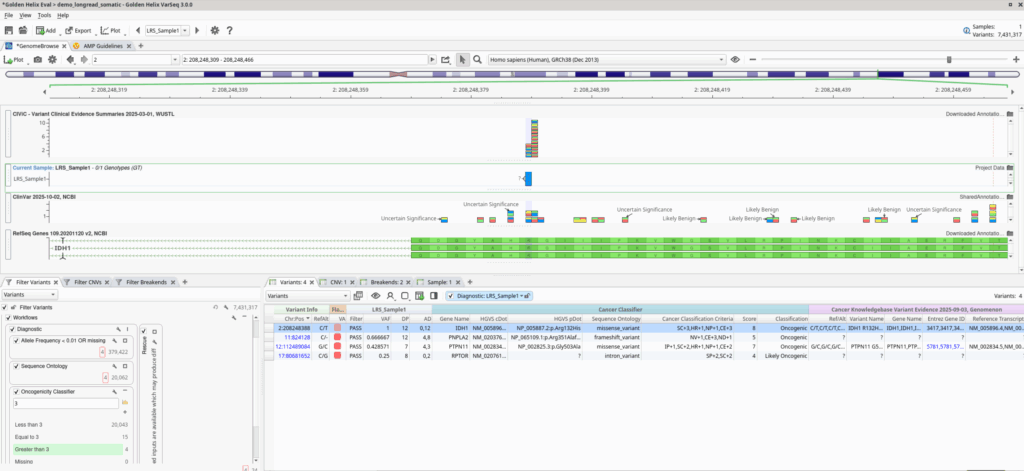
Why This Matters for Modern Somatic Testing
Somatic analysis often requires handling complex genomic alterations, large datasets, and frequent iterative updates to pipelines and interpretation guidelines. VSWarehouse 3.0 gives labs the ability to:
- Scale long-read somatic processing without reinventing infrastructure
- Maintain consistent best-practice workflows across users
- Capture complete audit histories of data, annotations, and reports
- Reduce manual overhead through automation and workspace-level versioning
As long-read sequencing expands into oncology, the ability to house both secondary and tertiary workflows under one system is becoming essential, and VSWarehouse 3.0 makes that possible.
Closing Thoughts
Golden Helix’s implementation of PacBio’s somatic workflow marks a significant step toward delivering end-to-end long-read analysis within a secure, centralized platform. Whether your team is exploring long-read applications or already scaling somatic pipelines, VSWarehouse provides the infrastructure needed to support accurate, repeatable, and automated genomic interpretation.



