About Darby Kammeraad
Darby Kammeraad is the Director of Field Application Services at Golden Helix, joining the team in April of 2017. Darby graduated in 2016 with a master’s degree in Plant Sciences from Montana State University, where he also received his bachelor’s degree in Plant Biotechnology. Darby works on customer support and training. When not in the office, Darby is learning how to play guitar, hunting, fishing, snowboarding, traveling or working on a new recipe in the kitchen.
Golden Helix has a long history of developing a high-quality NGS-based CNV caller. This has become even more apparent with our latest publication, where we teamed up with Twist Bioscience to demonstrate VarSeq capability of calling large copy number variants utilizing their clinical whole exome sequencing data. You can access the recent publication here: Analyzing Performance of Twist Bioscience Exome… Read more »
The release of VarSeq version 2.6.0 provides many new features. Most notably is our support for tertiary analysis of Pharmacogenomic (PGx) data. VarSeq not only calls the necessary gene diplotypes for your PGx panels but also handles large batches of samples from the called diplotypes to final report on drug recommendations. Here is a link to a recent webcast demonstrating… Read more »
Here at Golden Helix, we’ve sought to develop top-quality bioinformatic software to handle not only high-throughput clinical next-gen sequencing pipelines but also cater to research groups exploring their cohorts. Our tools fit in this NGS pipeline by importing the VCF and BAM files for downstream filtration, annotation, and classification/interpretation as well as clinical reporting. It was obvious early on and… Read more »
VarSeq serves as a streamlined approach to handle the rare variant analysis typically carried out for your next-generation sequencing data. Our team at Golden Helix seeks to simplify this process by automating the curation of staple databases needed for filtering and evaluation of clinically relevant variants. The goal we seek to accomplish with our software is to provide ease in… Read more »
If you stay current on the developments of Golden Helix features, you are aware of the substantial evolution of our copy number detection and evaluation capabilities in VarSeq. The process of CNV detection and evaluation is typically handled through the VarSeq graphic user interface. However, in some cases, users benefit from running this process via the command-line interface. Fortunately, Golden… Read more »
VCF file format comes with a lot of interesting quality assurance and statistics fields that can be used for filtering in VarSeq. Open your files in a text editor to see all the fields that are available in your files, each field will have a header line with a description of its content. See the VCF Specifications to help with… Read more »
In addition to Golden Helix providing easy-to-use genomic software, we also provide value to our users by automating the curation of public databases used in our tools. These annotation sources serve multiple purposes not only leveraging key fields in the filter chain but also automatically supplying evidence for the ACMG and AMP classifications reviewed in VSClinical. Supporting the curation of… Read more »
Clinical diagnostic efforts in next-generation sequencing are commonly defined at a gene panel level. The validation process of adding new genes to any diagnostic panel is ongoing, but labs typically construct and validate their clinical workflows for the current status of verified genes. This is not limited to primary finding results but can also include any incidental findings among the… Read more »
Next-gen sequencing (NGS) comprises many sophisticated steps that are often compressed into three major sections: library prep, sequencing, and data analysis. Obviously, the goal is to simplify each of these steps, but more often than not, there is a need for multiple tools to complete each one. Regarding the data analysis, Golden Helix seeks to provide simple yet comprehensive solutions… Read more »
One of the inherent realities of next-generation sequencing is the ongoing updates to the human reference genome—one of the strongest recommendations to take the original sequencing data and remap to the latest genome assembly. However, there are several reasons why remapping may be impractical. So, an alternative solution is needed to convert the data running through an initial mapping to… Read more »
In many cases, VarSeq users design their next-gen sequencing workflows for a clinical application. One of the major values of using VarSeq is the standardization of sample analysis via project templates for filtering down to rare variants and isolate any clinically relevant variant. However, VarSeq also doubles as a robust research application as well. There are specific algorithms that can… Read more »
Thank you to those who attended the recent webcast, “VSWarehouse: Tracking Changing Variant Evidence and Classifications”. For those who could not attend but wish to watch, here is a link to the recording. The webcast covered some general highlights of VSWarehouse value but also presented some specific capabilities covering the ClinVar classification tracker. Golden Helix provides complete solutions to handle… Read more »
In many cases, VarSeq users typically run single trio projects or perhaps an extended family project. Not only are all the inheritance model algorithms available in the VarSeq software to capture de novo, dominant, or recessively inherited variants but there are a number of quality control fields to help ensure the pedigree was set up properly. The last thing any… Read more »
Our latest release of the VarSeq software has had a major upgrade with the addition of the new CNV ACMG guidelines! Here are some recent webcasts we’ve given covering the new guideline tool: Family-Based Workflows in VarSeq and VSClinical A User’s Perspective: ACMG Guidelines for CNVs in VSClinical Not only does VarSeq 2.2.2 come with the new guideline tool, but… Read more »
VarSeq recently received major upgrades in a wide range of areas, one of these areas includes adding annotations such as GnomAD. This includes new fundamental methods of CNV ACMG guideline processing but also a large number of small additions in annotations. One addition is the application of gnomADs – Gene Constraints. This provides various metrics for pathogenicity on a per… Read more »
Our recent release of VarSeq 2.2.2 comes with a long list of upgrades and new features. In this blog post, we will demonstrate how defining sample phenotypes are available in VSClinical. One noticeable change is the ACMG guideline variant evaluation in VSClinical. Not only has this interface added CNV guideline evaluation, simplified the reporting process with embedded Microsoft Word and… Read more »
Our upcoming release of VarSeq is one of the largest we’ve ever had with our software! It comes with an extensive list of polishes and new features like our recently mentioned ACMG CNV classifier and a redesigned reporting interface with updated templates. Additionally, this new release is also paired with some major upgrades to our list of new and supported… Read more »
In this blog post, I will be analyzing a loss-of-function splice variant in MTHFR using VarSeq. In the search for clinically relevant variants contributing to rare disorders, efficient filtering strategies are an important step in eliminating disinteresting variants. However, any applied filters must also ensure no interesting variants inadvertently get filtered out. Golden Helix provides the tools to complete this… Read more »
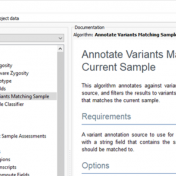
Those of you who have been attending our recent webcasts have learned about our upcoming VarSeq release. A part of that release will be an additional algorithm that will annotate variants matching the current sample. If you are not familiar with these webcasts, here are several on-demand webcasts I recommend to get you familiar with these new features: Evaluating Copy… Read more »
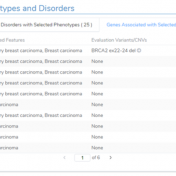
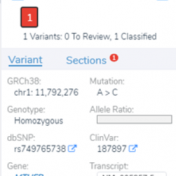
In a recent webcast, users were exposed to some new features upcoming in the next release of VarSeq. In this update, we will use an example de Novo CNV in a cardiomyopathy panel in VarSeq. There is a long list of new tools and polishes to the software but one major upgrade is the inclusion of the ACMG and ClinGen… Read more »