
Golden Helix’s VSClinical AMP guidelines platform, powered by our Golden Helix CancerKB has long been trusted by clinicians and researchers for generating report-ready clinical cancer biomarker interpretations. To further elevate the coverage and currency of our somatic variant evaluation capabilities, we’ve integrated Genomenon’s Cancer Knowledgebase (CKB) into both VarSeq and VSClinical AMP. This partnership represents a significant enhancement in somatic variant analysis, bringing global expertise and faster updates into the hands of our users. By combining Genomenon’s CKB with Golden Helix’s platform, we are expanding the breadth of curated evidence available for interpreting genetic variants in cancer, empowering clinical decision-making with greater speed and broader coverage.
Genomenon’s Cancer Knowledgebase (CKB) is one of the most extensive and rapidly updated databases of actionable cancer biomarkers. It provides over 48,000 variant summaries and curated evidence for over 2,100 genes, with rapid updates to ensure users always have access to the most current and clinically relevant data. This is particularly valuable in the ever-evolving field of cancer genomics, where discoveries and therapeutic options emerge at a rapid pace. Genomenon CKB integrates evidence from a wide range of international professional guidelines, offering a broader perspective on variant classification and clinical relevance. This comprehensive coverage ensures that users can rely on the most diverse and up-to-date resources for their somatic variant analysis, including cutting-edge insights from FDA-approved therapies, companion diagnostics, and emerging treatments.
Streamlining Variant Filtering with Genomenon CKB
The integration of Genomenon CKB into VarSeq brings powerful new tools for variant filtering. With CKB’s extensive evidence tracks for variants, regions, and fusions, users can easily identify actionable biomarkers linked to clinically relevant drugs and therapeutic strategies. Whether analyzing SNPs, indels, CNVs, or fusion events, Genomenon allows users to filter for variants associated with FDA-approved drugs, companion diagnostics, and emerging therapies, all based on AMP tier-level evidence. This comprehensive filtering process enables researchers and clinicians to focus on high-priority variants while navigating complex cases, such as Variants of Uncertain Significance (VUS) and fusions, making the interpretation process faster and more efficient.
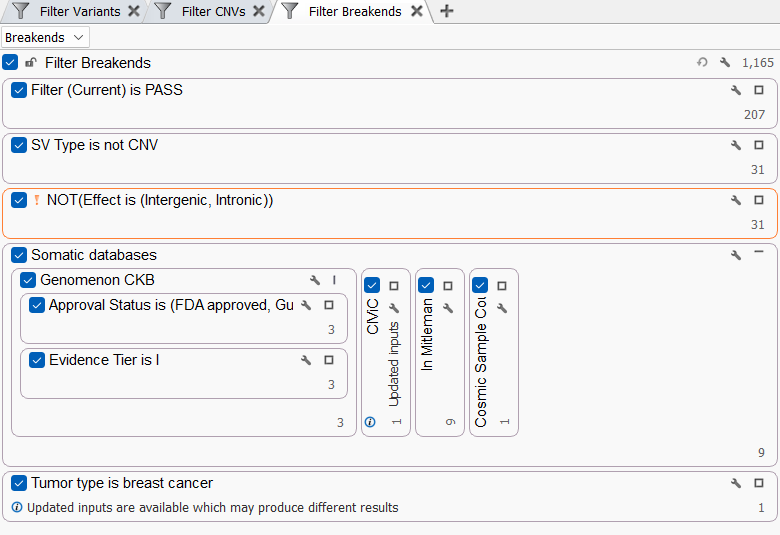
Integrating Genomenon CKB with VSClinical AMP for Report-Ready Interpretations
Once filtered, the curated evidence from Genomenon CKB is seamlessly integrated into VSClinical AMP, providing a unified workflow that supports AMP-based variant classification. VSClinical AMP is designed to generate report-ready interpretations by leveraging the robust data available through both Golden Helix’s curated sources and Genomenon’s CKB.
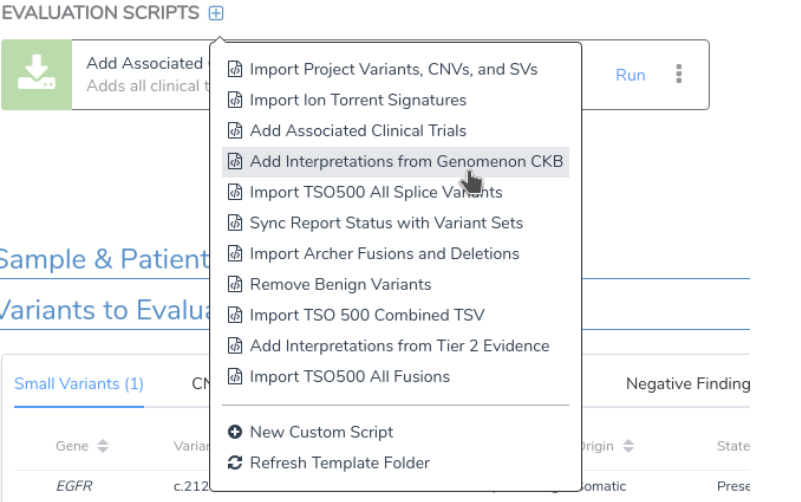
The integration of Genomenon’s Cancer Knowledgebase (CKB) into Golden Helix’s platform significantly expands the depth and scope of our somatic variant analysis tools. With faster updates, a global perspective, and comprehensive variant evidence, Genomenon enhances the speed and accuracy with which clinicians can interpret cancer variants and generate actionable insights.
By seamlessly incorporating Genomenon CKB into VarSeq and VSClinical AMP, we offer a powerful, integrated solution for clinical genomics. Whether for variant filtering, AMP classification, or report-ready interpretations, this integration supports clinicians in making more informed decisions based on the most current, relevant, and actionable evidence available.


