
In the dynamic field of pharmacogenomics (PGx), the human leukocyte antigen (HLA) system stands out as a critical factor in personalizing drug therapies, particularly in avoiding severe adverse reactions. However, as highlighted in a comprehensive review on the HLA system’s genetics and clinical testing, these genes are notoriously challenging to analyze due to their extreme polymorphism, complicating alignment and variant calling in next-generation sequencing (NGS) pipelines. Compounding this, current CPIC guidelines lack defined alleles for determining HLA diplotypes in NGS workflows, leaving a gap in standardized reporting.
Enter VarSeq—a versatile tool that empowers users to bridge this divide. By enabling the manual definition of diplotypes, VarSeq seamlessly integrates HLA insights into PGx reports, delivering actionable drug prescription recommendations even amid these hurdles. This capability not only enhances clinical decision-making but also paves the way for more robust, patient-centered drug recommendations.
VarSeq PGx Capability
Star Alleles are called from imported variant data in the VarSeq software and called with our PGx algorithm. The resulting star alleles and diplotypes are listed in the PGx genes table, which shows the predicted phenotype and drug recommendation details. You can see in Figure 1 an example output highlighted for CYP3A5 showing the called diplotype and alleles tested for their determination. Plotted in the image is the single core variant necessary to determine the star allele *3.
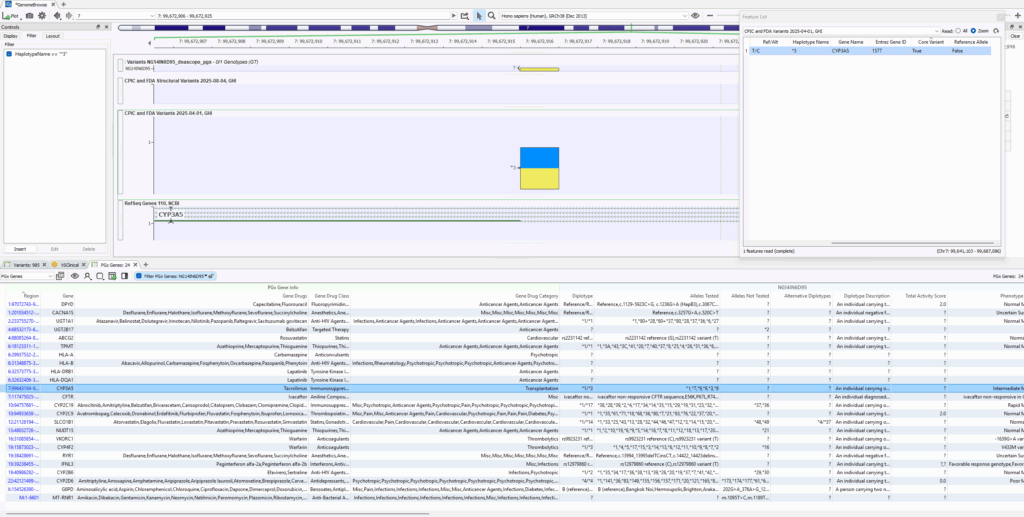
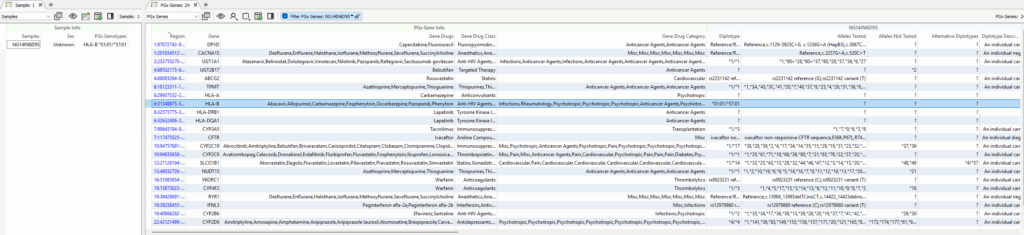
Now compare the output to HLA genes in Figure 2, where missing diplotypes and alleles are tested. Does VarSeq support processing HLA reportable outcomes? The answer is YES! VarSeq absolutely can handle these HLA diplotypes. We just have to overcome a limitation of the sourced CPIC database itself.
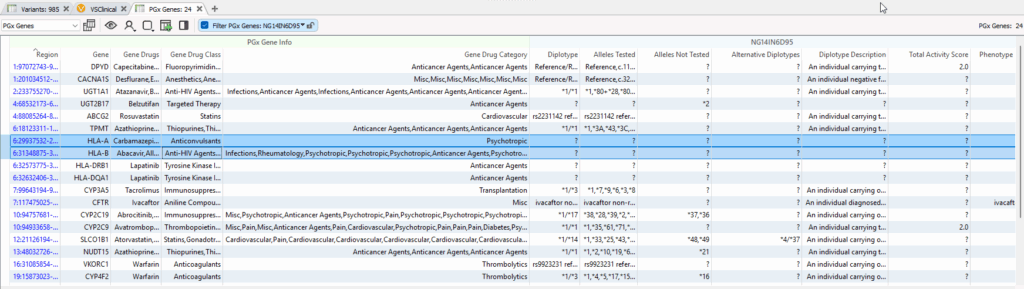
PGx Annotations
If you browse the CPIC guidelines site, you can access a breakdown of the expected variants contributing to each star allele, among other detailed guideline references. Figure 3 is a side-by-side comparison between HLA-B and CYP3A5. Notice the highlighted red box showing some supplemental information where users can download the allele definition table, and HLA-B does not list these alleles. Figure 4 also illustrates this limitation in PharmGKB/ClinPGx, where no specific variants are listed but the diplotypes themselves.
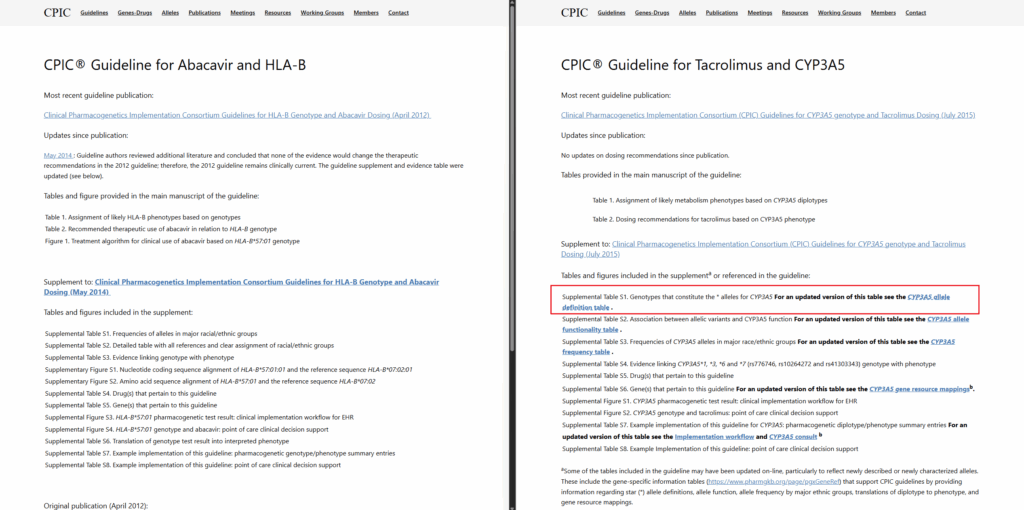
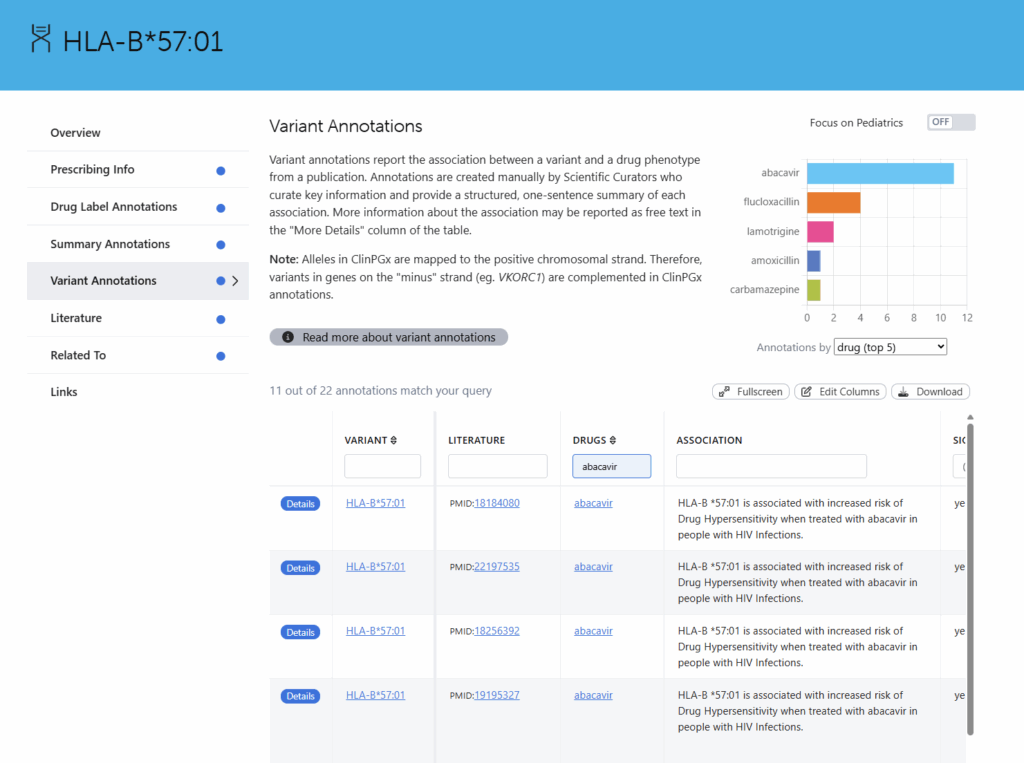
So, to account for these undefined variants, you can define a field for PGx Genotypes in Figure 5, the samples table in VarSeq, to include any other reportable diplotypes that may have individual variants missing from the databases. This now accounts for HLA-B in the PGx gene table.
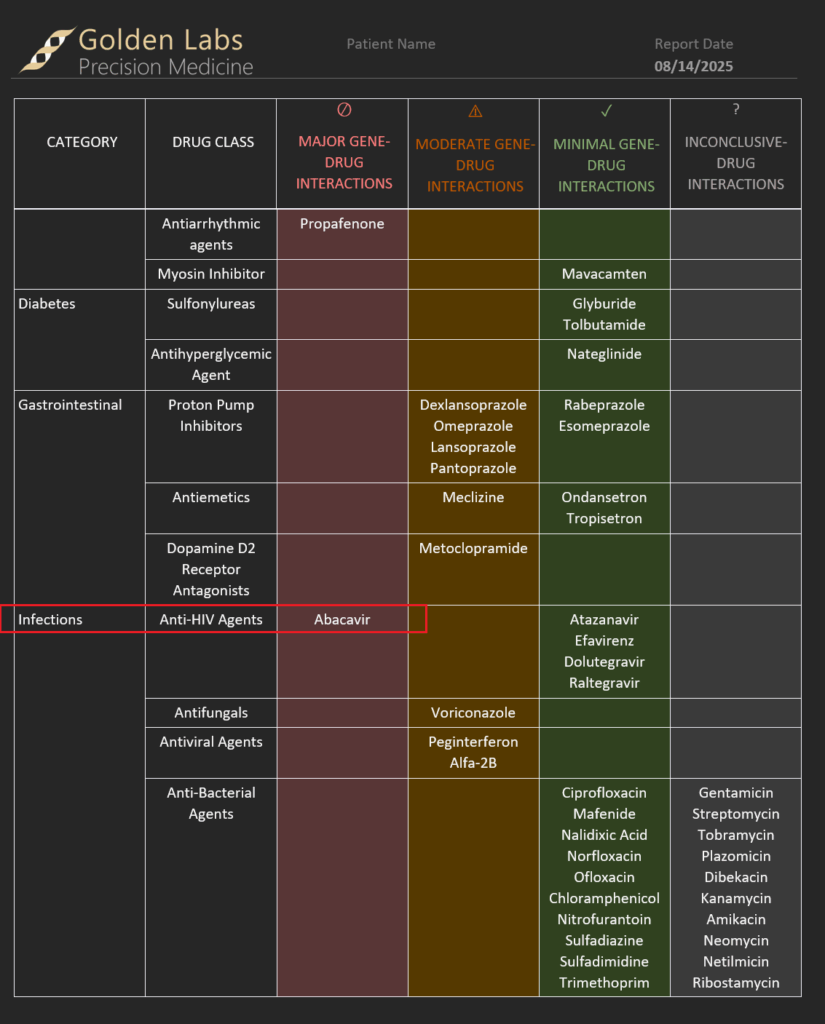
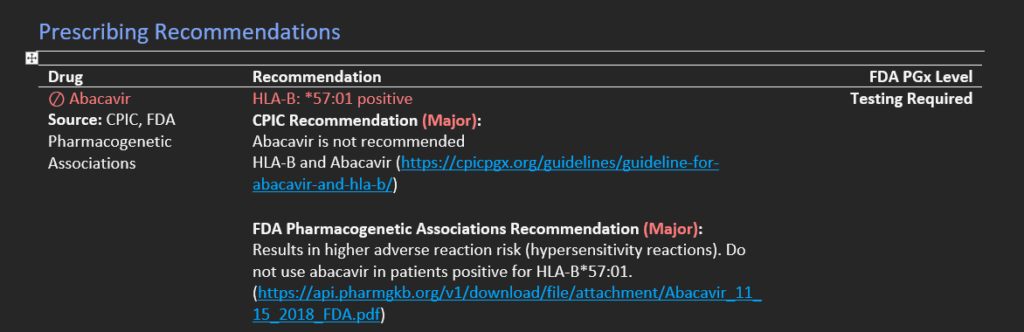
Moreover, these reportable outcomes will manifest in the report as well. Figure 6 shows some snapshots of the VarSeq report breaking down major, moderate, and minimal interactions that now account for HLA-B impact on Abacavir.
In conclusion, while the complexities of HLA genes result in some current limitations for seamless integration into NGS-based pharmacogenomics workflows, VarSeq emerges as a powerful solution that puts control back in the hands of users. By allowing manual definition of diplotypes where variant data is absent, VarSeq ensures that critical HLA insights are not overlooked, enabling comprehensive PGx reports complete with tailored drug recommendations. This flexibility not only addresses immediate gaps in standardized allele definitions but also advances the promise of precision medicine, helping clinicians mitigate risks like severe adverse reactions and optimize therapies for individual patients.
As the field evolves, VarSeq will be instrumental in turning genomic challenges into actionable opportunities for better healthcare outcomes.



