When performing variant classification using VarSeq’s ACMG classifier, a large number of variants inevitably fall into the category of variants of uncertain significance (VUS). This is because much of the evidence required for confident classification cannot be programmatically evaluated.
Take PS3, for example, the criterion for functional evidence. This criterion requires well-established in vitro or in vivo studies, but assessing study design, methodology, and quality of results demands expert review. Similarly, PS4, which considers case-control data, relies on variant frequency comparisons across affected and control populations. Such data are often scattered across multiple publications and rarely exist in a structured database format.
Without access to this deeper body of evidence, automated pipelines remain limited in their ability to resolve uncertain variants.
How Mastermind Fills the Gaps
This is where Genomenon Mastermind comes in. Mastermind provides a comprehensive catalog of pre-classified variants for germline analysis, including complete classifications and associated ACMG criteria for more than 150,000 variants. Each classification is curated by experts, ensuring that even evidence types requiring literature review and critical evaluation are incorporated. By leveraging this curated knowledgebase, laboratories can integrate expert-reviewed variant interpretations directly into their analysis workflows.
We are excited to announce the release of the Genomenon Mastermind Curated Variants annotation track. With this new annotation source, VarSeq users with a Genomenon license can:
- Directly integrate curated classifications into VarSeq filter chains.
- Automate the application of curated evidence in VSClinical’s ACMG Guidelines workflow.
- Streamline reporting, with curated classifications and inline references available for direct inclusion in clinical reports.
Incorporating Mastermind into the VarSeq Filter Chain
Let’s examine the tangible benefits of incorporating Genomenon Mastermind into an existing VarSeq filter chain. In our first example, we see the results of a standard filtering workflow that includes quality filters followed by a final classification filter:
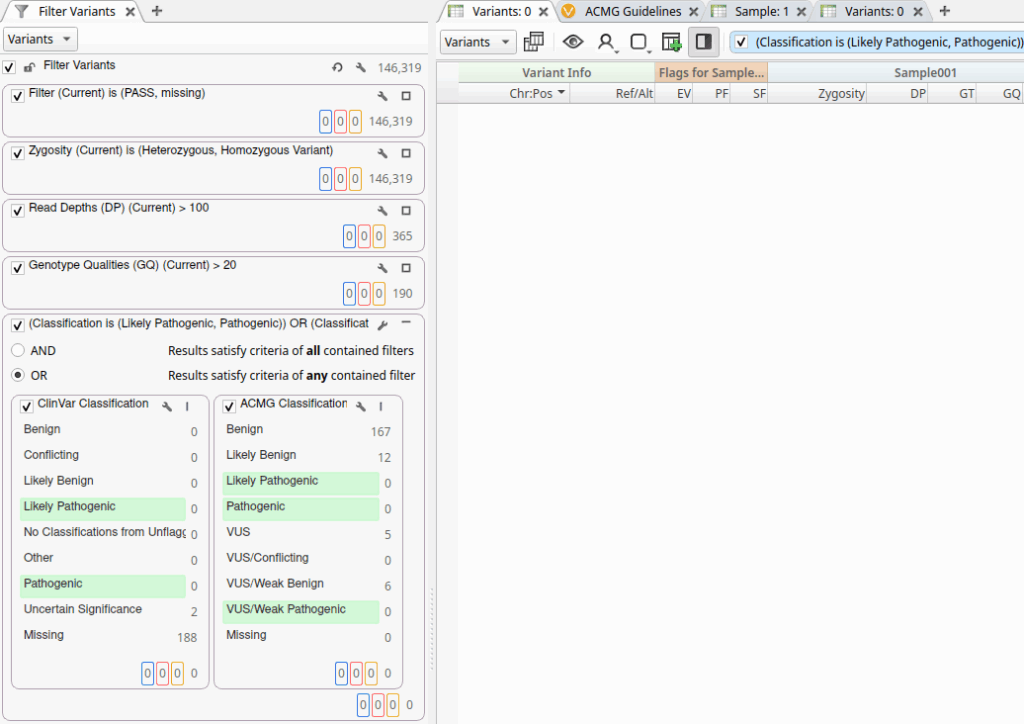
This classification filter incorporates classifications from both ClinVar and VarSeq’s ACMG Classifier algorithm, but in this particular sample, we don’t identify any variants with evidence of pathogenicity using these sources alone. However, by leveraging the new Genomenon Mastermind track, we can identify variants that may not be obviously pathogenic based on evidence available to VarSeq’s ACMG Classifier:
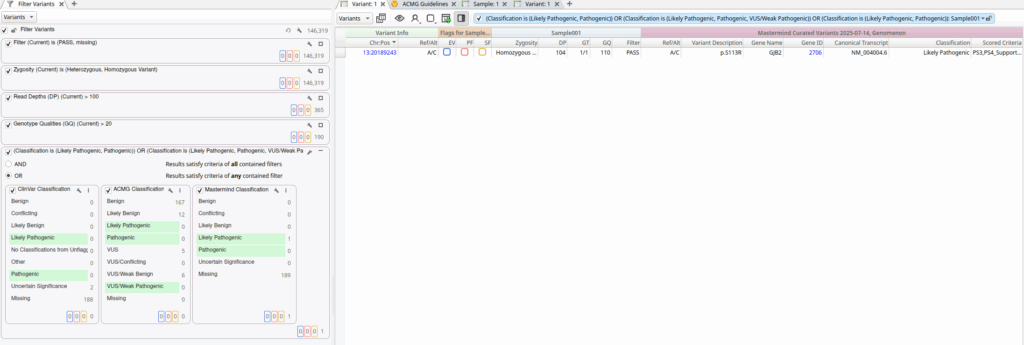
From the above filter chain result, we can see that our sample harbored a single variant that is classified by Genomenon as Likely Pathogenic that was not identified by either ClinVar or VarSeq’s ACMG Classifier. This demonstrates the critical value of Genomenon’s expert curation in identifying clinically significant variants.
Integration with VSClinical
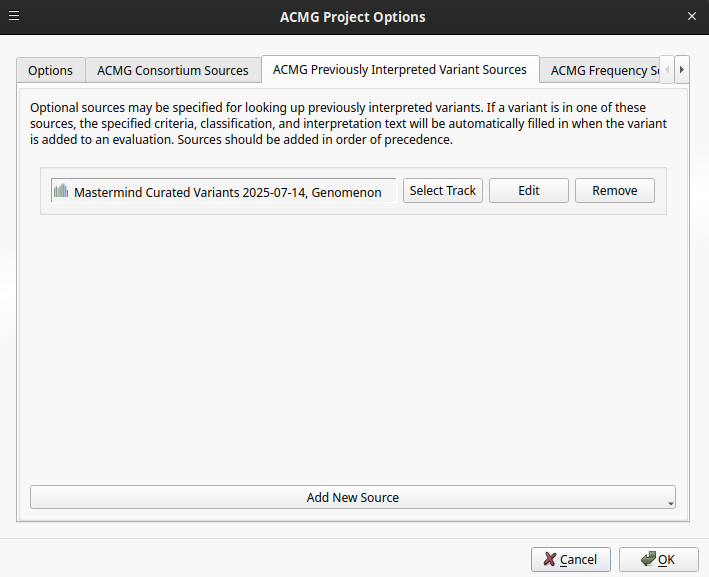
Let’s take a look at how Genomenon Mastermind can be integrated into the ACMG Guidelines workflow in VSClinical to automatically classify this variant and incorporate the evidence into a clinical report. By adding the Mastermind Curated Variants track as a Previously Interpreted Variants source, curated evidence is automatically applied to variants during evaluation.
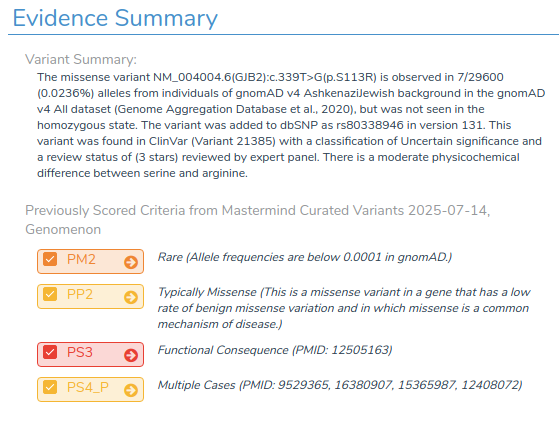
In this case, when evaluating the variant identified by our filter chain, we see that:
- The variant is automatically classified as Likely Pathogenic.
- All relevant ACMG criteria from Mastermind are applied.
- Supporting comments with inline references are automatically included.
These comments can be directly incorporated into the interpretation, which automatically generates an interactive list of inline citations supporting the classification. To explore the evidence, we can click any citation to view the complete abstract of the referenced publication.
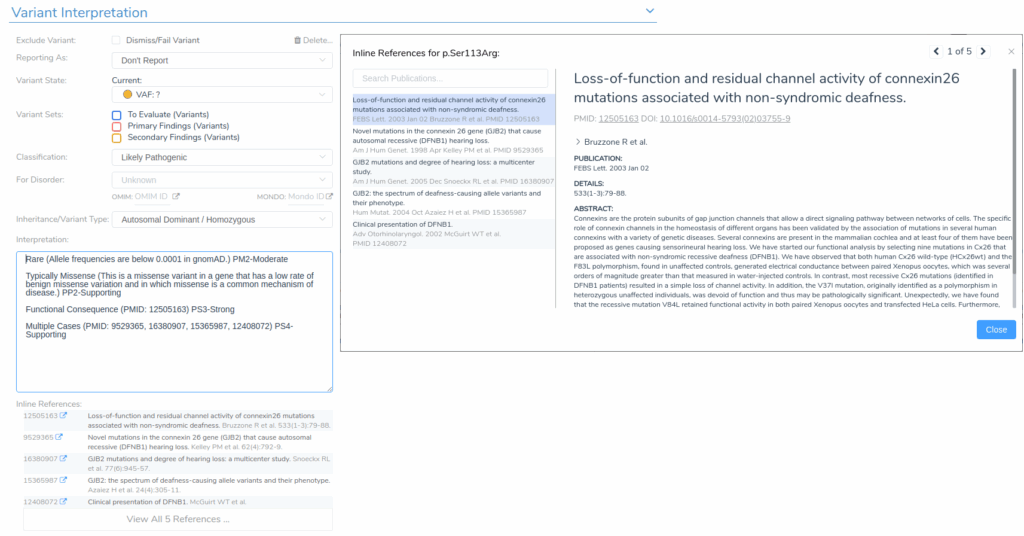
From there, we can finalize the interpretation, incorporating evidence from the relevant publications, which will be integrated into the automatically generated clinical report using VSClinical’s report generation system.
Conclusion
Variant interpretation is a complex process that often requires expert evaluation of literature and evidence beyond the reach of automated algorithms. With the release of the Genomenon Mastermind Curated Variants annotation track, VarSeq users can now close this gap by integrating expert curated classifications directly into their workflows. Please reach out to our team if you’re interested in learning more about how you can incorporate this powerful resource into your existing interpretation workflows.