End-to-End Structural Variant Analysis for Long-Read Data with VarSeq and VSWarehouse

Long-Read Data
Long-read sequencing technologies like PacBio HiFi have transformed our ability to detect structural variants (SVs) with greater accuracy and resolution than ever before. Unlike short-read sequencing, which often struggles to span the repetitive or complex genomic regions where SVs are most common, long reads can reliably resolve insertions, deletions, inversions, duplications, and translocations that would otherwise go undetected. At Golden Helix, we’ve built comprehensive support for PacBio long-read SV analysis directly into VSWarehouse and VarSeq, covering everything from raw variant calling to clinical interpretation and report generation.
Ready-to-Use PacBio Workflows in VSWarehouse
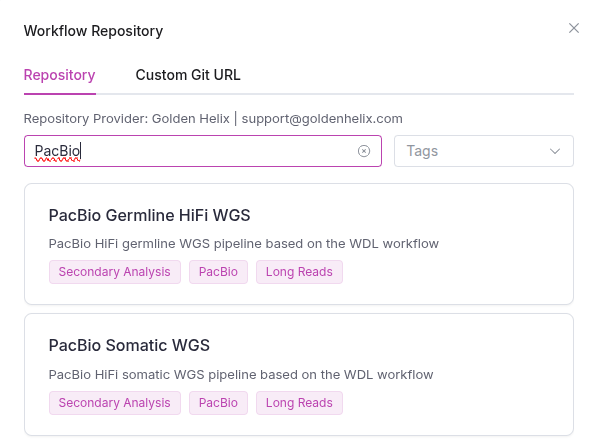
VSWarehouse includes two purpose-built workflows for PacBio whole-genome sequencing data, both available for download from our Workflow Repository:
- PacBio Germline HiFi WGS Workflow: designed for constitutional variant analysis, this workflow uses Sawfish to call structural variants alongside small variants and CNVs.
- PacBio Somatic WGS Workflow: designed for tumor/cancer analysis, this workflow leverages a combination of Severus, Wakhan, and CNVKit to provide a comprehensive picture of somatic structural variation.
Both workflows are fully automated and culminate in the generation of a VarSeq project with structural variants, small variants, and CNVs already imported and ready for analysis.
Annotating and Filtering Structural Variants
Once a workflow completes, the resulting VarSeq project comes pre-configured with a set of annotations and filters tailored to structural variant interpretation. These templates are completely customizable, giving laboratories complete control over the annotation tracks, filter thresholds, and variant categories to match their specific clinical or research requirements.
The included annotations include gene information and effect predictions, allowing users to immediately distinguish between clinically distinct consequences, such as in-frame fusions or transcript ablations.
The screenshot below shows the filtered set of SVs called by Sawfish in our PacBio Germline HiFi workflow, illustrating how users can quickly identify a small list of potentially clinically relevant structural variants:
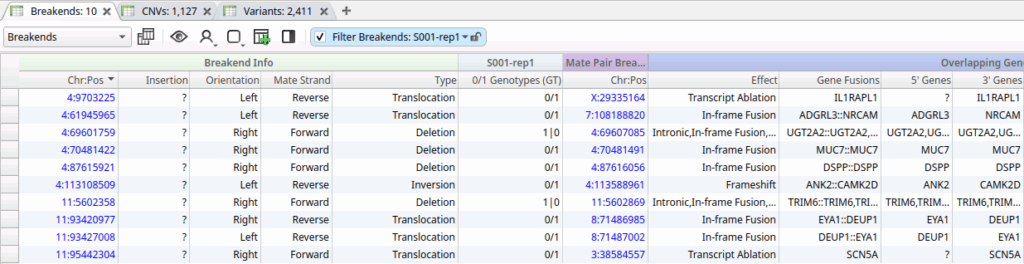
Clinical Interpretation in VSClinical
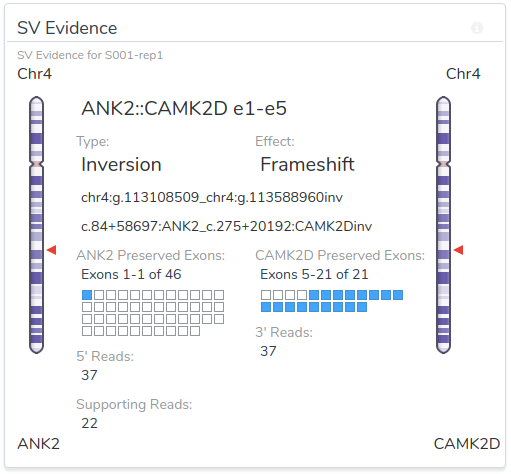
Once filtered, structural variants can be directly imported into VSClinical for interpretation and classification. VSClinical provides a rich set of tools specifically designed to support SV review, including variant summary information and breakpoint visualization.
The VSClinical visualization makes it clear that this chromosome 4 inversion disrupts the 5′ coding region of ANK2. The breakpoint occurs after exon 1, producing an ANK2::CAMK2D fusion transcript that is predicted to introduce a frameshift early in the ANK2 coding sequence.
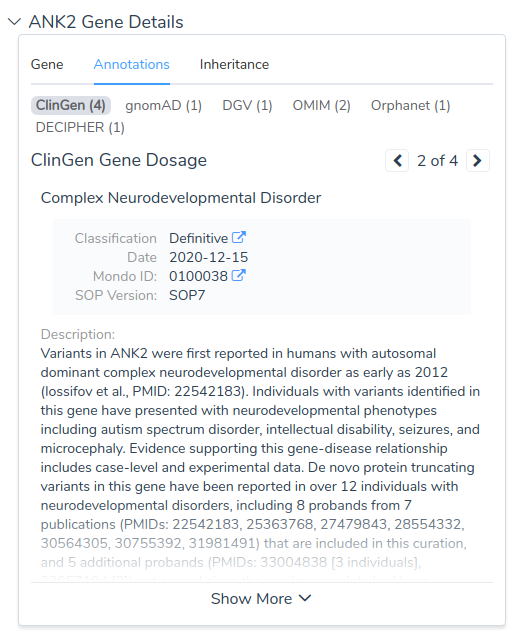
VSClinical also integrates information from resource like ClinGen and OMIM including detailed references with links to PubMed, streamlining the process of gathering the supporting evidence required to draft variant interpretations.
Once structural variants have been interpreted and classified, they can be incorporated into a finalized clinical report using VSClinical’s customizable report templates. Reports are generated automatically based on the variants and interpretations captured during the review process, ensuring consistency and saving time.
Conclusion
As long-read sequencing continues to expand in both research and clinical settings, having a scalable and customizable analysis framework is essential. With VSWarehouse and VarSeq, laboratories can confidently leverage long read data to uncover clinically meaningful structural variants and bring clarity to even the most complex genomic rearrangements. Please reach out to us if you would like to learn more about support for long-read analysis in VarSeq and VSWarehouse.