
Carrier screening is an important component of reproductive genetic testing, helping identify individuals or couples at increased risk of passing on inherited conditions to their children. When both partners carry a pathogenic or likely pathogenic variant in the same gene associated with an autosomal recessive condition, or when a pathogenic variant is identified on an X-linked gene, there may be an elevated risk for offspring to be affected.
VSClinical provides a streamlined, guided workflow for evaluating and reporting carrier status results, integrating variant interpretation, evidence curation, and report generation into a single, intuitive interface. In this post, we’ll walk through how VSClinical can be used to perform Carrier Analysis incorporating paired sample data, highlighting how to configure the evaluation, interpret findings, and generate a clear, clinician-ready report.
Setting Up the Carrier Analysis Evaluation
Before starting a new carrier analysis evaluation, it’s essential to configure VSClinical to properly handle data from both partners. First, verify that the Evaluation Type is set to Partner and that the correct sample has been selected as the partner sample. This configuration ensures that clinically relevant variants from both the primary sample and the partner are included in the analysis and covered by the final clinical report.
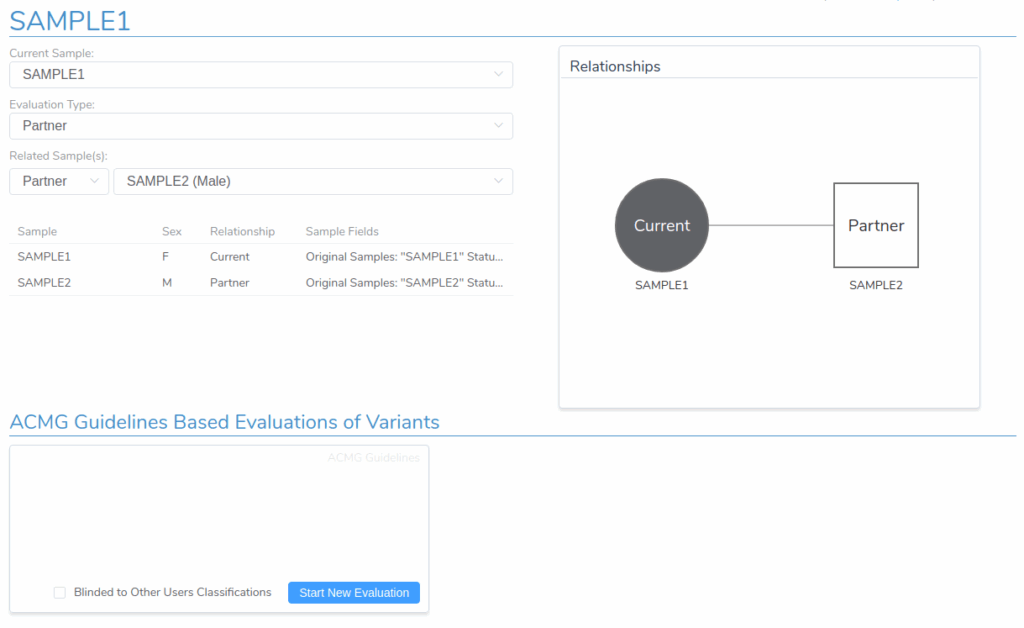
Adding Filtered Variants from Both Samples
Once the evaluation has been created, you can add filtered variants for both samples using the Add Variants From Project dialog. First, click Prepare to Add to select the filtered variants from the primary sample. Then switch to the partner sample by clicking the arrow icon at the top of the dialog. After selecting Prepare to Add again, check Score All Recommended Criteria to automatically apply classification criteria across both samples. Finally, click Add 3 Variants to include them in the evaluation.
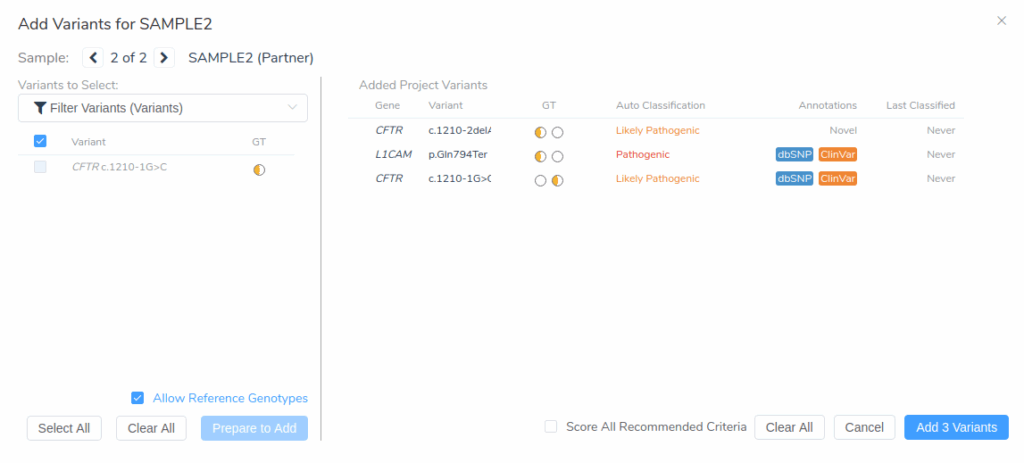
The Variants to Evaluate section now lists all selected variants across the two samples.
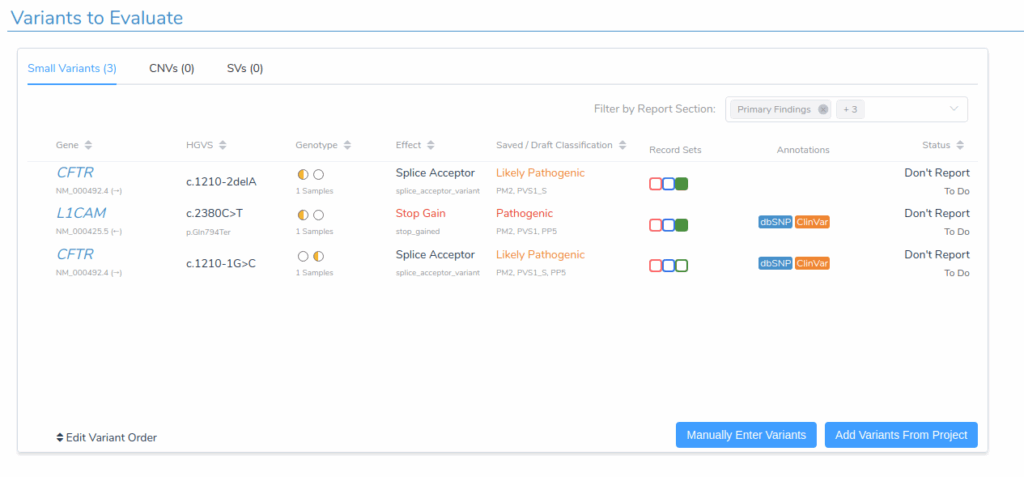
Reviewing Variant Evidence and Classification
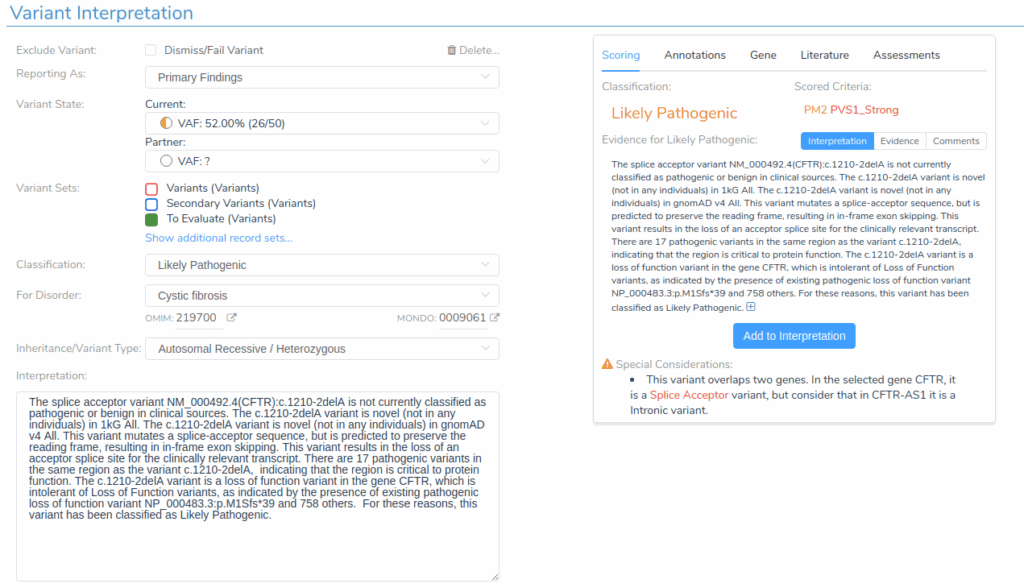
Switching to the Variants tab reveals comprehensive information supporting each variant’s classification. VSClinical provides detailed evidence summaries that facilitate efficient variant interpretation in accordance with the ACMG Guidelines.
The Evidence Summary section displays the recommended classification and the specific criteria used as evidence. For example, the CFTR c.1210-2delG variant shows a classification of Likely Pathogenic, which is supported by the criteria PM2 and PVS1_S.
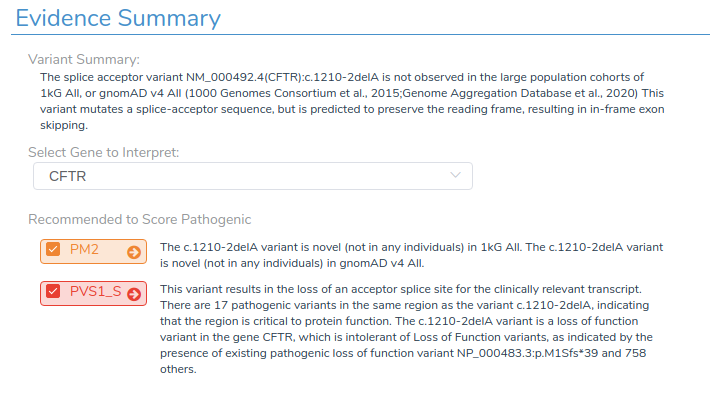
For each variant, we review the applicable criteria, assign the appropriate report section, and finalize the interpretation.
Analyzing Gene-Level Inheritance Patterns
Once variant interpretation is complete for both samples, the Genes tab provides valuable summary information for all genes containing primary or secondary findings. This gene-level view is particularly important in carrier analysis, as it can be used to reveal inheritance patterns that determine reproductive risk.
In this evaluation, both the primary sample and the partner have a heterozygous pathogenic variant in the CFTR gene. When viewed together, this indicates a compound heterozygous inheritance pattern, meaning the couple is at increased risk for having a child affected with cystic fibrosis.
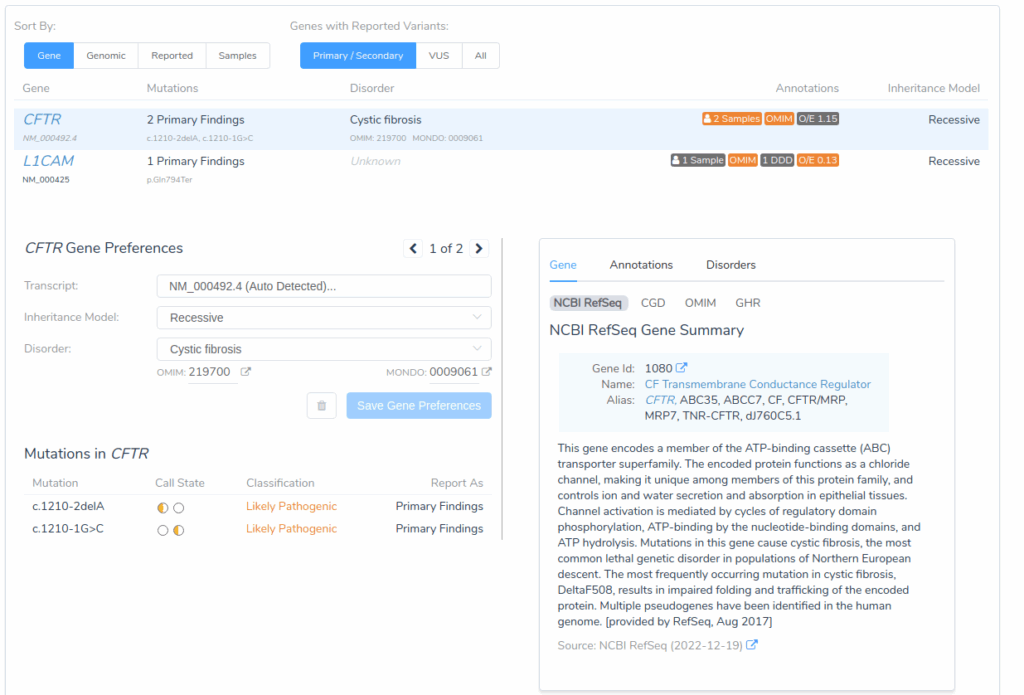
This gene-level view helps clinicians quickly identify which genes pose reproductive risks based on shared carrier status between partners.
Generating the Clinical Report
With variant interpretation and gene analysis complete, we move to the Report tab to generate the final clinical report. For this demonstration, we’re using the default Carrier Report Template that ships with VarSeq, though the system supports customizable templates to meet specific institutional requirements.

The generated report consolidates information from both the primary and partner samples, clearly indicating whether variants were detected in genes associated with specific disorders for which carrier status is clinically significant.
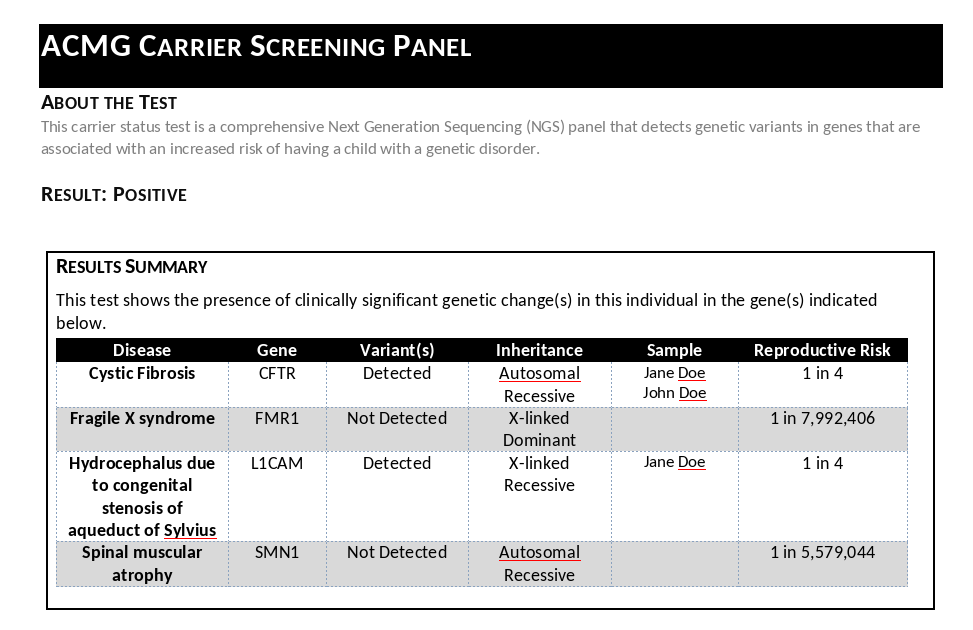
The report reveals that variants were detected in two genes: CFTR and L1CAM. This finding indicates increased reproductive risk for two conditions:
- Cystic Fibrosis (associated with CFTR variants)
- Hydrocephalus (associated with L1CAM variants)
The report format facilitates clear communication with patients and informs providers about the specific reproductive risks identified through carrier screening.
Conclusion
VSClinical simplifies the complex process of carrier status evaluation by integrating variant analysis, evidence scoring, and clinical reporting within a single guided workflow. By pairing results from both individuals in a couple, laboratories can quickly identify shared variants in genes associated with inherited disorders and generate comprehensive, professional reports that support informed reproductive decision-making.
Whether your lab performs high-throughput carrier screening or targeted variant interpretation, VSClinical provides the flexibility and automation needed to ensure consistent, accurate, and clinically meaningful results. Please reach out to our team if you would like to know more about how VarSeq and VSClinical can be used to streamline your carrier analysis workflow.



