Recently, Golden Helix Field Application Scientists Rana Smalling and Andrew Legan led a webcast on Secondary NGS Workflow Automation in VSWarehouse. The presentation walked through the process of turning FASTQ data into completed VarSeq projects and clinical reports with just a click of a button.
During the Q&A, a few key topics were raised, and we’ll discuss those further in this blog post. These questions revolved around workflow customization, pharmacogenomic analysis, and managing compute resources in VSWarehouse.
Workflow Customization
“How can the user modify automated workflows if they already have custom scripts”
Maybe your lab already uses custom bioinformatics scripts in its secondary analysis pipeline, or perhaps you’re a bioinformatician keen to develop custom scripts in the future. A core feature of VSWarehouse automation is that it simplifies complex bioinformatics tools so that the user doesn’t need to edit code. For bioinformatics-inclined users, the platform provides the tools needed to write their own scripts.
Workflows in VSWarehouse are written in YAML, making them approachable and easily modifiable. The integrated VSCode editor supports syntax highlighting and inline documentation, allowing you to efficiently edit scripts and immediately test-run workflows. Users can wrap their own scripts or swap in custom tasks as stages of an existing workflow.
If editing a script seems daunting, know that training on task and workflow customizations is included in the support provided by our FAS team during VSWarehouse installation, setup, and user onboarding.
Pharmacogenomic Genotyping
“There was a CypCall step included in the short read PGx workflow. Is that used for the PacBio long read workflow as well?”
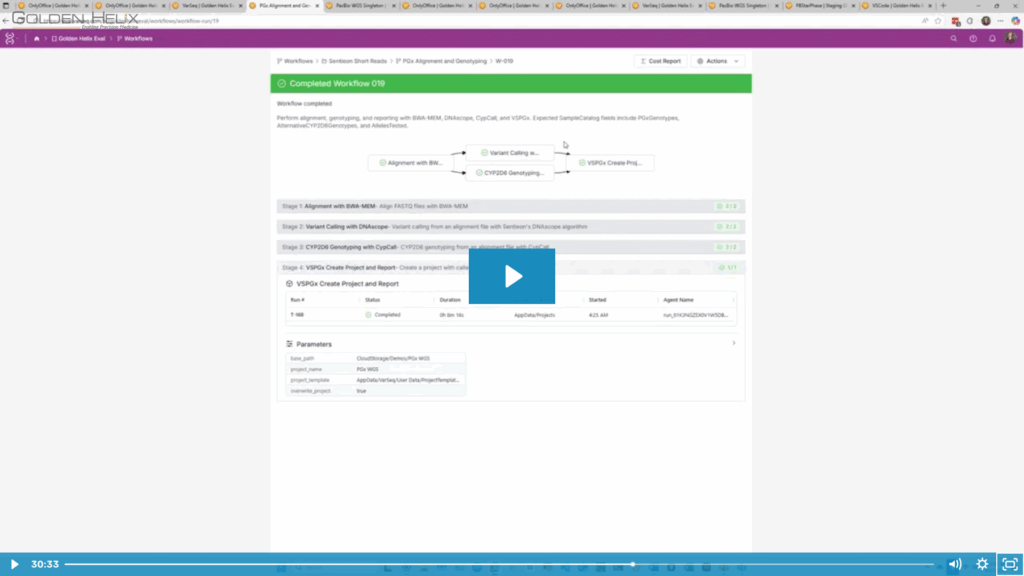
CypCall is a competitive option for labs seeking CYP2D6 star-allele genotyping. It pairs well with short-read secondary workflows, like Sentieon DNAscope, and is already integrated into short-read alignment and variant calling workflows that ship with VSWarehouse.
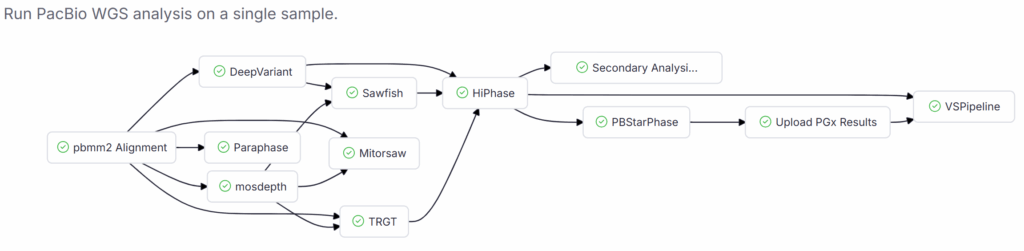
For long-read workflows, for example, with PacBio HiFi data, pharmacogenomic interpretation is handled by PBStarPhase, which integrates outputs from DeepVariant, Paraphase, TRGT, Mitorsaw, Sawfish, and Mosdepth. Our PacBio WGS workflow adds these star alleles to the workspace Sample Catalog and generates a full PGx report covering drug interactions, evidence levels, and clinically significant alleles. Golden Helix is officially PacBio-Compatible, and this workflow is just one illustration of that partnership.
As with all Golden Helix software, VSWarehouse is constantly updated to expand support, and we have workflows in development for alignment and variant calling with ONT EPI2ME, DRAGEN, Archer VariantPlex, and other bioinformatics pipelines. Regardless of which platform you prefer, VSWarehouse is a stellar option for automation of PGx analysis and reporting.
Scalability and Resource Management
“Can multiple workflows be run in parallel, and how can we control the compute power that is allocated to each workflow?”
Scalability is another central theme of automation in VSWarehouse. The platform supports parallel execution of workflows, as well as simultaneous execution of stages within workflows.
Admins can configure machine allocation in the Admin Center. For example, for cloud accounts, admins can set the maximum number of instances per agent profile to fine-tune scaling. If all resources are in use, VSWarehouse will queue tasks until the required resources become available. That way, labs can maximize throughput while staying within the provided quotas. Each workflow allows real-time monitoring of cloud compute costs, generating a concise expense report after each run.
You don’t need a cloud account to utilize the flexibility of VSWarehouse’s compute architecture. On-premises deployments can also configure the resources allocated to the modular compute structure of locally run workflows.
We are excited to have new and existing customers test out the automated workflows now available in VarSeq Warehouse 3.0! Please contact us at [email protected] if you would like to learn firsthand how our tools can facilitate your NGS analyses.