Golden Helix GenomeBrowse: Interactive Genomic Visualization
Powerful genome browser software for visualizing BAM, VCF, BED, and annotation tracks. Available as a free standalone application and built directly into VarSeq for synchronized variant analysis.
A genome browser software is an essential tool for any bioinformatician or clinical researcher, providing a visual window into the complex landscape of the human genome. By transforming raw sequencing data into intuitive, interactive maps, these tools allow users to validate variant calls, inspect coverage, and understand the genomic context of their findings.
While many browsers exist, the challenge lies in performance and clarity. Visualizing massive BAM and VCF files requires significant computational efficiency to ensure fluid navigation. Golden Helix GenomeBrowse addresses these needs as a high-performance desktop application, delivering stunning visualizations that outperform web-based alternatives.
Use Golden Helix GenomeBrowse as a free standalone genome browser for research and academic work, or experience its full potential built into VarSeq, where it becomes a context-aware visualization tool synchronized to your variant analysis workflow.
Two Ways to Use Golden Helix GenomeBrowse
Golden Helix GenomeBrowse is available as a free standalone desktop application for any researcher, and is also the built-in visualization engine powering VarSeq clinical workflows.
Golden Helix GenomeBrowse Standalone
A high-performance desktop genome browser for visualizing BAM, VCF, BED, and annotation tracks. Free for academic use, with support for any species and genome assembly.
- Open BAM, VCF, BED, CRAM, FASTA, and WIG files directly
- Overlay ClinVar, gnomAD, gene tracks, and custom annotations
- Multi-species support with custom genome assemblies
- Export publication-quality plots for papers and presentations
Golden Helix GenomeBrowse in VarSeq
Inside VarSeq, Golden Helix GenomeBrowse becomes a synchronized visualization tool, automatically displaying the genomic context for whatever variant, CNV, or region you're analyzing.
- Synchronized selection: click any variant, CNV, break-end, or coverage region and Golden Helix GenomeBrowse navigates instantly
- Sample-aware BAM display: automatically loads and plots the BAM file for the sample you're working with
- Automatic trio plotting: in family analysis, displays the proband's BAM alongside both parents' BAMs for de novo variant validation
- Tumor-normal pairing: in somatic analysis, automatically plots the tumor BAM and its matched normal for side-by-side comparison
What to Look for in a Genome Browser
Native Performance
Avoid the lag of web-based tools. A native desktop application utilizes your hardware's full power for smooth data interaction.
Broad Format Support
Ensure support for standard formats like BAM, VCF, FASTA, BED, WIG, and CRAM without complex data conversion.
Integrated Annotations
Direct access to public databases (ClinVar, gnomAD) and gene tracks without leaving the browser interface.
Publication Exports
The ability to export high-resolution, print-ready visuals for journal submissions and clinical documentation.
Tertiary Integration
The browser should connect directly to your variant interpretation platform for instant validation of filtered results.
Multi-Species Support
Full support for any genome assembly, including human, plant, and animal builds. Easily add custom species and annotations for diverse model organisms.
Synchronized Visualization Inside VarSeq
When Golden Helix GenomeBrowse runs inside VarSeq, it transforms from a standalone viewer into a context-aware visualization tool. Every action in VarSeq (selecting a variant, navigating a CNV call, inspecting a structural break-end) automatically updates the view to show the relevant genomic region with the right sample data.
Click-to-Visualize
Select any variant, CNV, break-end, or coverage region in VarSeq and Golden Helix GenomeBrowse instantly navigates to that position, displaying the raw alignment data and annotation context you need to validate the call.
Automatic Trio BAM Plotting
In family analysis, Golden Helix GenomeBrowse automatically loads the proband's BAM alongside both parents' alignments. Instantly see whether a variant is de novo, inherited from a carrier parent, or a compound heterozygous hit.
Tumor-Normal Side-by-Side
For somatic variant analysis, Golden Helix GenomeBrowse automatically pairs the tumor BAM with its matched normal control, making it easy to confirm somatic-only calls and identify artifacts from germline variants.
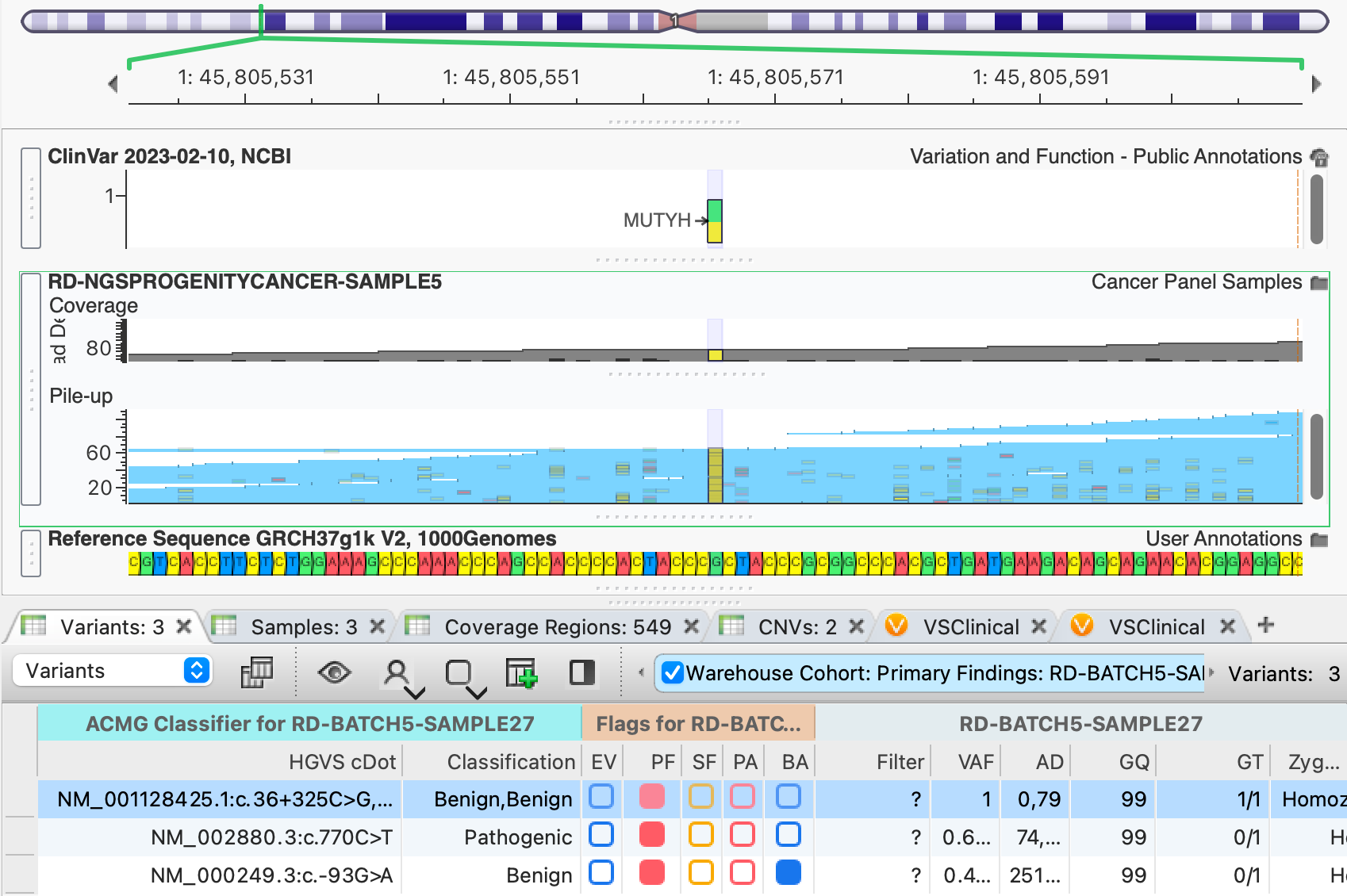
"Click a variant in VarSeq and Golden Helix GenomeBrowse jumps to that position with the sample's BAM and annotations already loaded."
Integrated Visualization Workflow
Direct Data Load
Open BAM, VCF, BED, or CRAM files directly from local storage or remote servers without complex conversion steps.
Coordinate Navigation
Jump to specific genes, regions, or variant coordinates with instant rendering and fluid pan/zoom controls.
Visual Inspection
Overlay ClinVar, gene tracks, and sample alignments to validate findings and identify biological context.
Clinical Export
Export publication-quality plots directly for journal submissions or clinical reports with cross-plot anchors.
Visualize Every Layer of Genomic Variation
From small SNVs to large-scale structural variants, Golden Helix GenomeBrowse provides specialized views for every data type.
- Alignment Pileups: Identify coverage gaps and alignment artifacts in BAM files.
- CNV Regions: Visualize read-depth changes and structural breakpoints at scale.
- Clinical Context: Overlay ClinVar, gnomAD, and OMIM annotations directly.
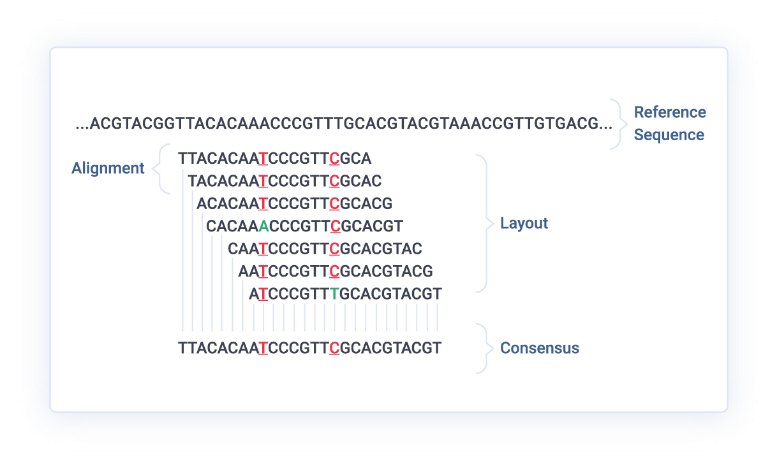
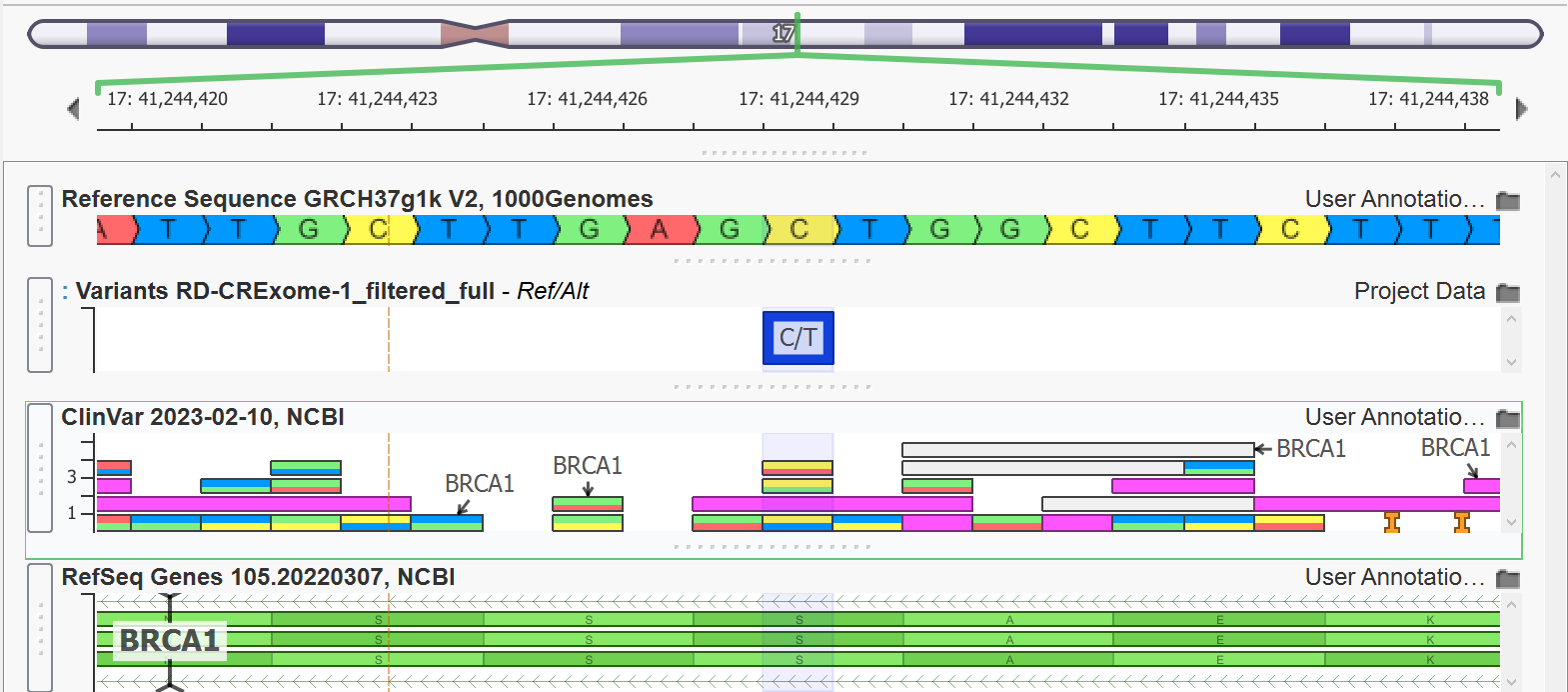
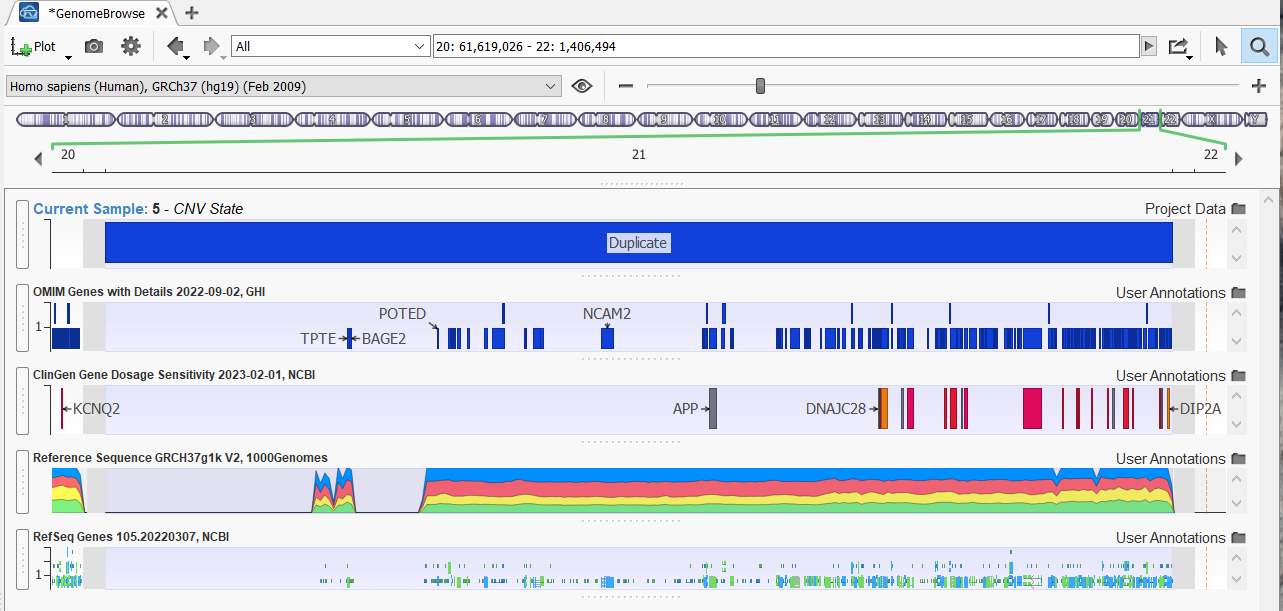
Specialized Use Cases
Variant Validation
Confirm variant calls by inspecting the raw alignment data. Identify artifacts that automated callers might misinterpret, such as strand bias or homopolymer issues.
Variant Analysis WorkflowStructural Variation
Visualize CNVs and large-scale rearrangements across multiple samples. Synchronize plots to identify de novo events or inherited deletions in trio analysis.
CNV Analysis DetailsAcademic Discovery
A free genome browser for academia that supports discovery across any species. Create publication-quality plots for research papers and presentations.
Genome Analysis SolutionsFrequently Asked Questions
Is Golden Helix GenomeBrowse really free?
Yes. GenomeBrowse is available without charge for academic use. Commercial laboratories and diagnostic centers should contact Golden Helix for commercial licensing terms.
What operating systems are supported?
Golden Helix GenomeBrowse is a native desktop application available for Windows 10+, macOS 10.13+, and major Linux distributions including Ubuntu 18.04/20.04 and CentOS 7.
Can I visualize non-human species?
Absolutely. Golden Helix GenomeBrowse supports multiple genomic builds beyond humans, including plants and animals. You can easily add custom species with their own reference sequences and annotations.
How does Golden Helix GenomeBrowse handle data privacy?
Since it is a native desktop application, all data is processed locally on your computer. Your sensitive alignment and variant files never need to be uploaded to the cloud, ensuring complete data sovereignty.
How does Golden Helix GenomeBrowse work inside VarSeq?
GenomeBrowse is the built-in visualization engine for VarSeq. When you select any variant, CNV, break-end, or coverage region in VarSeq, GenomeBrowse automatically navigates to that position and loads the sample-specific BAM file. In trio workflows, it automatically displays the proband alongside both parents' BAMs; in somatic workflows, it pairs tumor and matched normal BAMs side-by-side.
Visualization Insights & Webcasts
Learn advanced visualization techniques for variant validation, structural variation analysis, and publication-ready plotting.
Ready to See Your Data with Base-Pair Clarity?
Download Golden Helix GenomeBrowse for free or explore how it powers synchronized visualization inside VarSeq.