In our SVS 8.6.0 release, we updated our Annotate and Filter Variants feature to utilize our powerful VarSeq annotations. Annotations can be run against gene, interval, variant, and tabular tracks, including RefSeq, ClinVar, CADD, OMIM and OncoMD. The new streamlined dialog allows users to select track specific options and to set up custom filters.
While our public annotation repository has been previously available in SVS, they are now more powerfully integrated with more advanced filter and annotation control.
New in this release is our licensed premium annotation sources including CADD pathogenicity scores, full OMIM gene and variant annotations and the curated cancer mutation, gene and drug targeting mutation data from OncoMD.
Fast Annotations & Powerful Filters with SVS
We have previously written about how the VarSeq transcript annotation algorithm tries to build on the best from Annovar, VEP and snpEff. Now, that algorithm will be the default in SVS for annotating against a gene source and will provide the same useful and detailed fields, including the selection of clinically relevant transcripts.
To streamline the common annotation and filtering process, we have rebuilt SVS’s “Annotate and Filter” feature as a wizard where you can specify options on which annotation outputs you want as well as what filters you may want to perform on your data for each selected annotation source.
You will also have options related to how SVS will annotate your variants on a source-specific basis.
With the RefSeq Genes 105v2 NCBI source, for example, you can specify the splice site boundaries used to determine splice-site variants or to only annotate verified mRNA transcripts (the default).
In addition, filters can be set up to subset down the original dataset into just the variants you are interested in, such as selecting variants with a Missense or Loss of Function change to the transcript selected as Clinically Relevant.
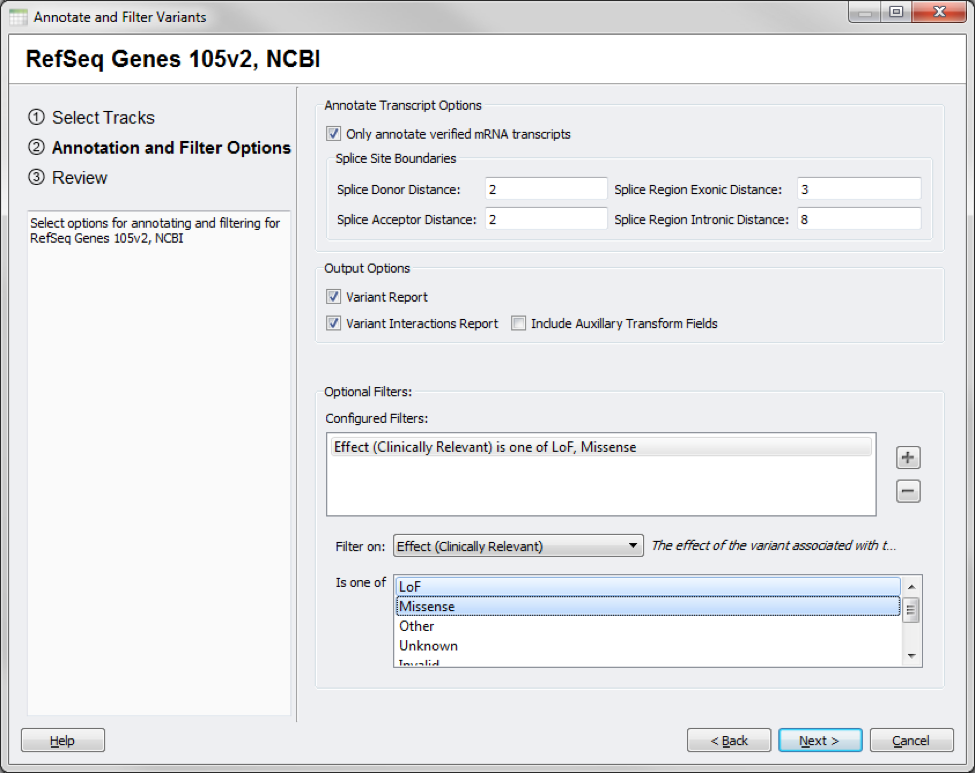
Annotating against RefSeq filtering on clinically relevant effects
You can also select multiple annotations sources at a time and use their annotation output or configure additional filters.
Say that along with a gene-based filter, you want to also narrow your variants to those that are Pathogenic or Likely Pathogenic variants according to ClinVar!
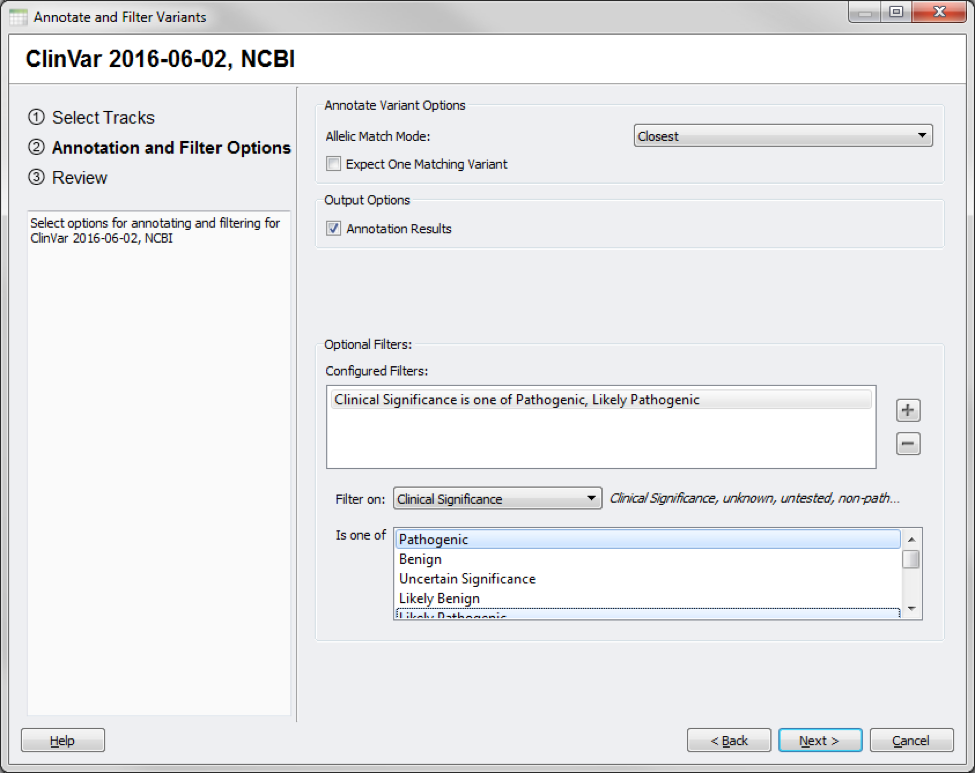
Annotating against ClinVar filtering on Clinical Significance
In fact, any field in these annotation sources can now be used to configure custom filters that can be applied alongside the process of producing the annotation results.
This includes common use cases like filtering rare variants from ExAC, filtering functional variants from the dbNSFP voting algorithm and even our new licensed annotation sources.
After the annotation process is complete, a filtering results spreadsheet will display showing whether or not a variant passed each of the configured filters as well as a combined “Is Filtered?” field, then the original spreadsheet will be filtered down to just the passing variants.
The new annotation features will also be able to utilize our secure annotation sources, such as CADD Scores, OMIM and OncoMD.
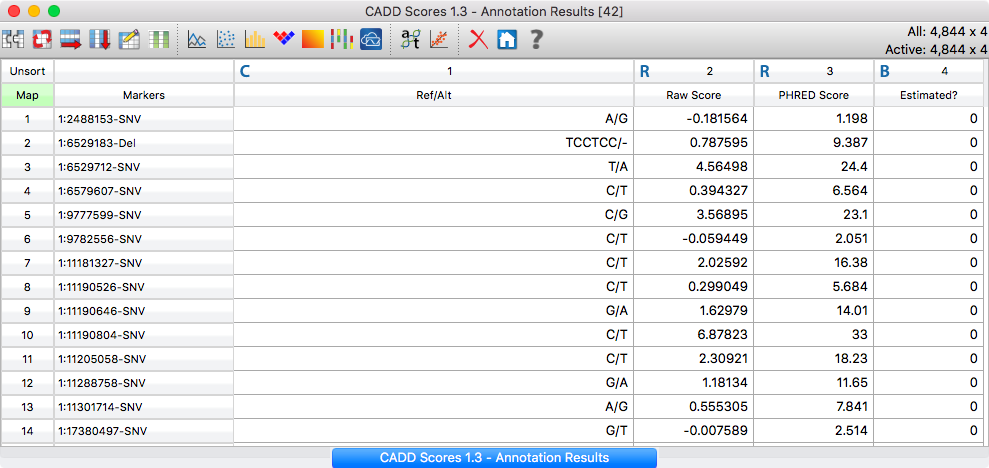
CADD Scores Annotation Results
If you are interested in accessing these new annotation options for your SVS workflows, please reach out to your Golden Helix area director.
We believe that this improvement to our Annotate and Filter Variants Wizard will help you streamline your analysis workflow by accessing our robust annotation algorithms from VarSeq and customizable filtering options. We are looking forward to the next SVS release!
