Using the AMP Guidelines for Rhabdoid Tumor Analysis in VSClinical
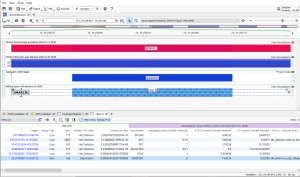
Introduction: Malignant Rhabdoid tumors (MRT) are among the most aggressive and lethal forms of infant and child cancer (1). These tumors are characterized by an unusual combination of mixed cellular elements similar to but not typical of teratomas and can originate at any anatomic location. When MRTs are present in…
Read more →