Targeted Gene Panel Analysis for Clinical Diagnostics
From hereditary cancer and cardiac panels to oncology profiling and prenatal screening, VarSeq delivers end-to-end panel analysis with configurable filter workflows, ACMG/AMP classification, and regulatory-ready clinical reporting.
Why Targeted Gene Panels?
Gene panels sequence a defined set of genes associated with specific conditions—delivering deep coverage of clinically relevant regions at lower cost and faster turnaround than exome or genome sequencing. A typical panel test produces over 2,000 variants per sample, and VarSeq's configurable filter workflows reduce that to single-digit clinically actionable findings.
Deep, Focused Coverage
Sequencing reads concentrated on target regions deliver 300x+ mean depth, enabling confident variant calling and limiting incidental findings to clinically relevant genes.
Faster Time to Report
Smaller data footprint means faster secondary analysis and fewer variants to review. What takes hours with genome data takes minutes with a focused panel.
Simpler Validation
Focused regions simplify analytical validation for laboratory and test accreditation. VarSeq's locked, versioned workflows and built-in coverage QC support reproducible, audit-ready processes.
Configurable Filter Chains That Narrow to What Matters
VarSeq's filter chain systematically reduces thousands of variants to a short list of clinically significant candidates. Each filter card is adjustable, saveable, and reusable across samples — here's an example of a typical germline panel workflow.
Zygosity Filter
Retain only heterozygous and homozygous variant calls, removing reference and no-call entries. Reduces 1,202 variants to 836.
Genotype Quality
Apply a quality threshold (GQ ≥ 30) tuned during validation to balance sensitivity and specificity, narrowing to 802 variants.
Warehouse Cohort
Filter against your lab's internal allele frequency database from VSWarehouse. Remove common variants seen across your cohort, reducing to 713.
Clinical Classification
Filter by ClinVar classification for Pathogenic and Likely Pathogenic variants, producing a final list of 3 variants ready for ACMG interpretation and clinical reporting.
Panels for Every Clinical Specialty
VarSeq supports the full range of targeted gene panels—from germline hereditary testing to somatic tumor profiling and prenatal screening—each with guideline-driven classification and clinical reporting.
Hereditary Disease Panels
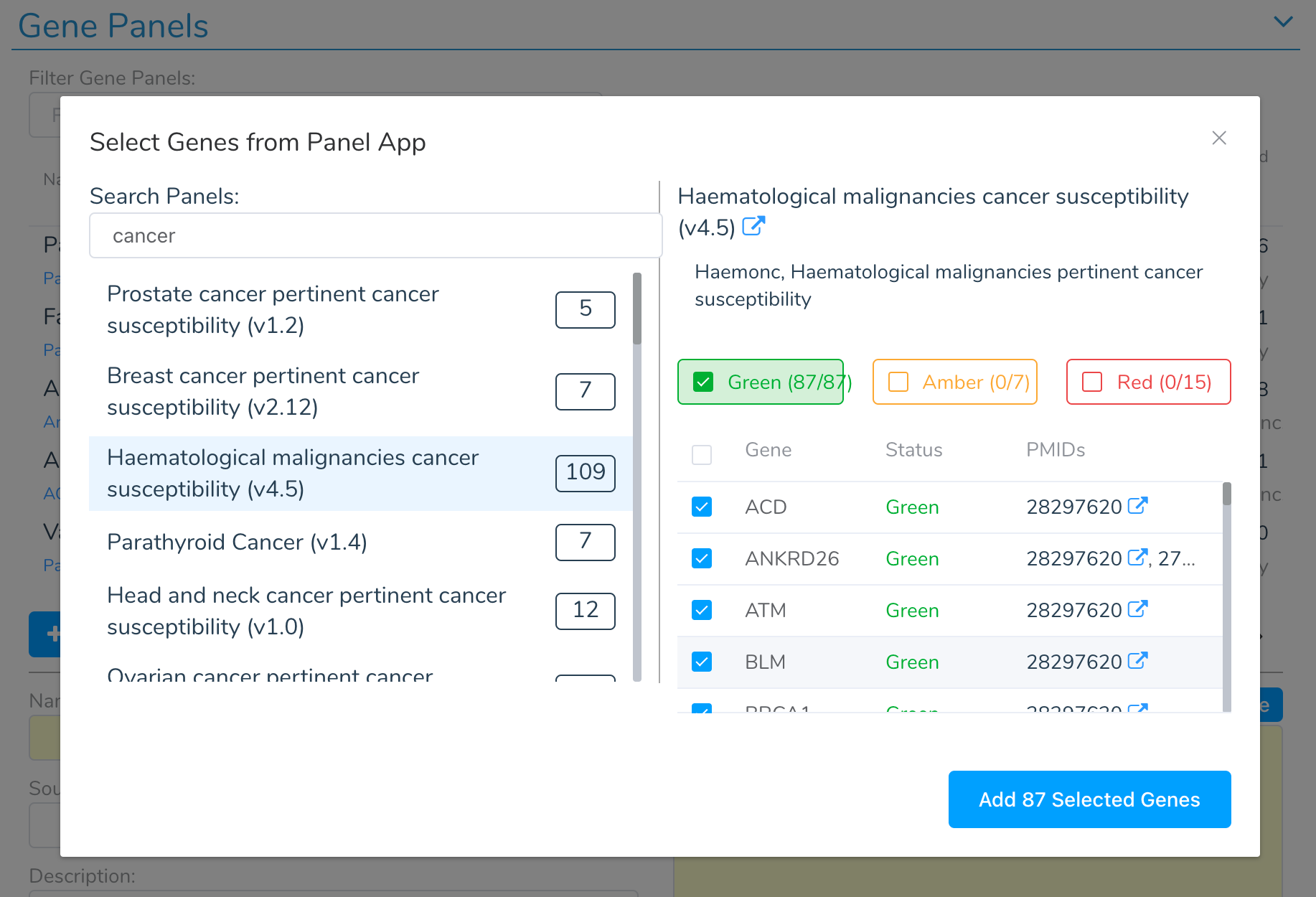
Hereditary cancer panels (BRCA1/2, Lynch syndrome, Li-Fraumeni), cardiac panels (cardiomyopathy, arrhythmia, aortopathy), and neurological panels (epilepsy, neurodevelopmental disorders) are interpreted using the ACMG/AMP germline classification framework with automated scoring of all 28 ACMG criteria.
Oncology Profiling Panels
Comprehensive genomic profiling panels like the TruSight Oncology 500 are fully supported with the AMP/ASCO/CAP somatic classification framework. VSClinical automates evidence aggregation, oncogenicity scoring, and tier classification across all biomarker types.
Missense, nonsense, and frameshift variants with oncogenicity scoring
Copy number amplifications, deletions, and gene fusions like BCR-ABL1
Tumor mutational burden and microsatellite instability for immunotherapy eligibility
Match biomarkers to NCI-funded trials by location, phase, and eligibility criteria
Prenatal & Carrier Screening Panels
Approximately 3% of deliveries are complicated by genetic or birth defects, and significant chromosomal abnormalities affect up to 1 in 140 live births. VarSeq supports prenatal gene panels and carrier screening with specialized workflows for monogenic disorders, CNV detection, and PhoRank phenotype-driven variant ranking.
Stop gained — loss of function due to protein truncation. ClinVar confirmed, population novel.
Frameshift — hereditary breast/ovarian cancer susceptibility. Secondary finding per ACMG list.
Heterozygous deletion — familial adenomatous polyposis. Detected via VS-CNV from panel depths.
Panel QC & Accreditation Readiness
CAP accreditation requires documentation of all algorithms, software versions, databases, and SOPs used in your bioinformatics pipeline. VarSeq's locked, versioned workflows and built-in QC tools meet these requirements out of the box.
BED File Region Filtering
Automatically restrict analysis to validated panel regions. Off-target variants are removed to maintain focus on primary findings and prevent false discoveries.
Coverage Verification & Must-Call Sites
Identify regions with sub-optimal read depth. Verify that critical hotspot sites have sufficient coverage, and flag failed regions as inconclusive in the report.
Versioned Pipeline & Audit Trail
Every analysis is reproducible with locked software versions, annotation databases, and filter configurations. Exception logs and upgrade tracking built in for CAP audit readiness.
The Complete Panel Analysis Toolkit
VarSeq provides a comprehensive ecosystem for targeted gene panel testing in clinical laboratories.
CNV Detection
Integrated clinical-grade CNV calling from panel read depths. Resolve single exon events.
Exome Analysis
Expand to whole exome sequencing when broader coverage is needed for unresolved cases.
GenomeBrowse
High-performance visualization of BAM and VCF files with interactive annotation overlays.
Whole Genome
Move to full-scale whole genome sequencing analysis for the broadest diagnostic reach.
Panel Analysis Insights & Webcasts
Stay updated with the latest in targeted gene panel testing, ACMG guidelines, and clinical implementation.
Ready to Streamline Your Panel Workflows?
Join the hundreds of clinical laboratories worldwide that trust Golden Helix for their targeted gene panel analysis.