Allow me to introduce you to Blaine Bettinger. Blaine is a patent attorney who holds a PhD in Biochemistry with a concentration in genetics. He is also a family history enthusiast who writes the Genetic Genealogist blog, where he gives commentary on applications of genomic science for advancing personal and family history research. I first learned about Blaine last May when he announced that he was committing his genomic data to the public domain and challenged any interested parties to analyze it and tell him what they learned. As a scientist, he was aware of the limitations of the technology, but was curious to know if anybody might find something surprising. As an intellectual property lawyer, he expressed interest in the question of whether it was even possible to truly commit one’s genomic data to the public domain. (Anybody who attended ICHG/ASHG in Montreal may have heard some of the debate about whether genomic research data belongs to the researcher, the study subject, the funding agency, or some other entity.) I immediately downloaded the data with grand visions of what I might do with it, but set it aside due to other obligations and forgot about it until rediscovering it just a few weeks ago.
I contacted Blaine to ask him about his experience since posting his data. He replied that he has yet to receive any real response to his challenge–a result which we both found very interesting in itself. Why hasn’t he received a response? Perhaps the reason is that the field of personal genomics is still in embryonic stages, and there just isn’t very much that we can do with SNP genotype data from a single genome. Indeed, Blaine has already published analysis results very similar to everything that I considered doing with his data. He posted information about his autosomal admixture from multiple tests, suggesting that he is between 84% and 98% European depending on the method used. He posted his mitochondrial haplogroup (Native American maternal lineage) and Y haplogroup (continental European paternal lineage). He posted Promethease reports that give insight into his susceptibility to a variety of traits based on published research. For example, he may have reduced risk of rheumatoid arthritis, but increased risk of type-2 diabetes. To me, family history research and determining disease risk seem like the most probable reasons for a person to pursue direct-to-consumer (DTC) genotyping services, and Blaine has covered those topics fairly well. I was fascinated by the information he posted and decided to take a little bit of time to understand his results and see what else could be learned from his genotypes.
The data Blaine posted consists of genotypes from four whole-genome SNP arrays which he ordered from two DTC genotyping companies. There were two arrays from Family Tree DNA, which appear to be based on the Affymetrix Axiom and the Illumina OmniExpress platforms. He also has the 23andMe v2 (customized Illumina 550) and v3 (customized Illumina OmniExpress) arrays. For the work I will show below, I used all of these except the 23andMe v2 product. This was my first experience with data generated by DTC genetics labs, and I found the quality to be very good. The call rates were high, and the agreement between the 3 platforms I examined was excellent. Perhaps I am too easily impressed, but I was very excited to discover that the genotypes from all three arrays had been resolved to agree with the human genome reference strand. Anybody who has ever tried to combine genotype data from multiple sources knows what a challenge it can be to deal with various strand definitions. In this case, it was a relatively simple exercise to merge Blaine’s multiple sources of genotype data and combine it with data from Phase 1 of the 1000 Genomes Project, allowing me to compare his genome with 1094 subjects from 14 reference populations.
My initial goal when I started working with Blaine’s data was to validate the information found in his 23andMe Ancestry Painting — a karyotype illustration with haplotypes colored to indicate continental origin. We frequently speak about admixture in statistical genetics, and I think that we sometimes focus so much on the population-level analysis that we forget what admixture looks like on an individual level. It is always refreshing to see admixture illustrated so clearly. The ancestry painting shows that Blaine’s genome is mostly European in origin, but he has several haplotypes originating in Asia and Africa — perhaps not surprising given that he can trace his maternal lineage to Central America. My results seem to agree with 23andMe. Figure 1 shows a plot of the first two principal components calculated using genome-wide SNP data from Blaine and 1,094 subjects from the 1000 Genomes Project.
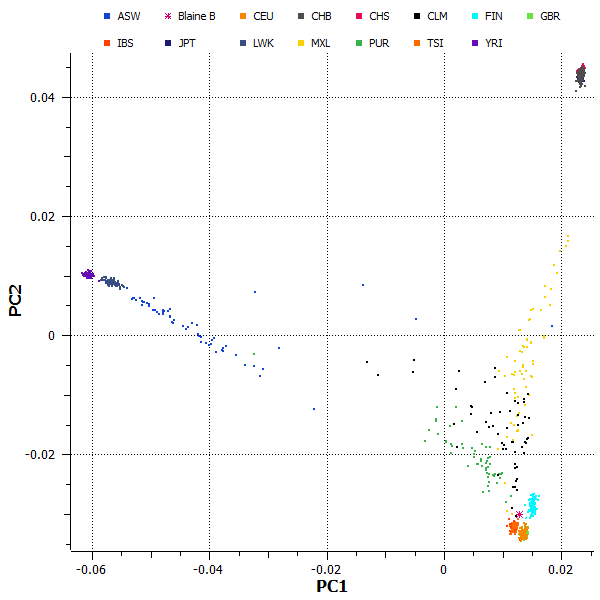
Figure 1: Principal components analysis of whole-genome data shows that Blaine Bettinger’s genome (marked with magenta asterisk) is very similar to reference populations from Europe.
We can see in this figure that Blaine’s genome is very similar to the European reference populations, as I expected based on his published admixture results. Although his genome is primarily European in origin, the ancestry painting tells us that Blaine should have an African haplotype near the q-terminus of chromosome 12. To confirm this, I repeated the PCA test using only the SNPs in the final 24 Mbp of chromosome 12. The results, as illustrated in Figure 2, indeed show that Blaine has an apparent mixture of African and European alleles in this area.
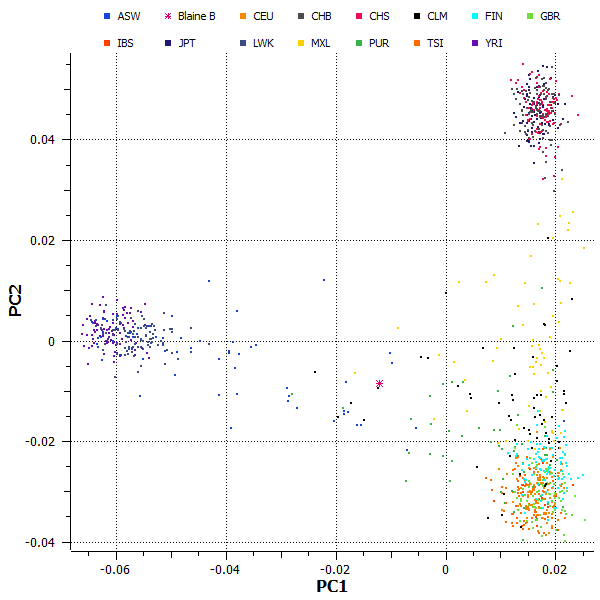
Figure 2: Principal components analysis of SNPs from a 24 Mbp region near the q-terminus of chromosome 12 indicates that Blaine (magenta asterisk) has an apparent mix of European and African alleles in this region.
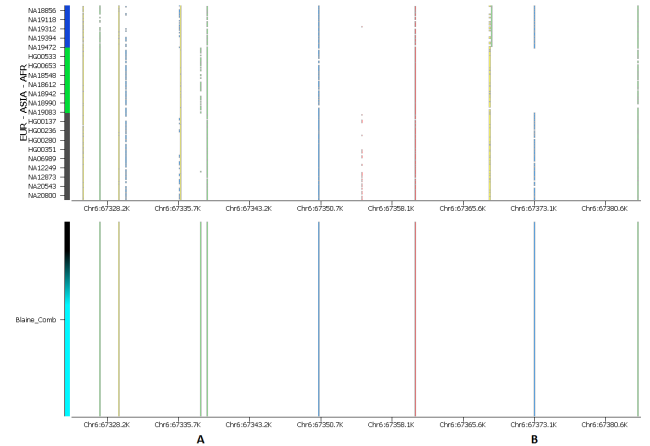
I followed a similar process to validate the large Asian haplotypes indicated by 23andMe on chromosomes 2, 6 and 13. The results in these areas were not as clear as the African result on chromosome 12, but each test grouped Blaine together with subjects from admixed American populations in places like Puerto Rico, Colombia and Mexico. A closer inspection of the data at the genotype level, as shown in Figure 3, gives more support to 23andMe’s determination for an Asian haplotype on chromosome 6.
Figure 3: SVS variant map comparing Blaine Bettinger’s genome to subjects from the 1000 genomes project. The upper frame shows data for 852 subjects from African (top), Asian (middle) and European (bottom) population groups, while the lower frame shows variant sites in Blaine’s data. 1000 Genomes Project variant sites are shown only if they were assayed on one of the three consumer arrays used in the present assessment of Blaine’s data. Note that at position A, Blaine has a variant that is found almost exclusively in Asian populations, while he has a variant at position B which is entirely absent in Asian populations. He is heterozygous at both positions, supporting the hypothesis that he carries both European and Asian haplotypes at this locus.
Here we can see that Blaine has a sequence variant that is entirely absent in the Asian reference subjects located very close to a variant that is observed almost exclusively in Asians. Perhaps unsurprisingly, he is heterozygous for both SNPs. There are many examples of this phenomenon nearby, and with higher-density data we would certainly see even more examples of this.
As I explore Blaine Bettinger’s genome, I am reminded of a conversation I had with statistical genetics colleagues at a recent conference. They said that they were involved in a GWAS project that included large numbers of young African-American subjects, and the analysis was severely complicated and confounded by admixture. Admixture, of course, is expected in African Americans, but they said that this project was unlike anything they had ever encountered due to high levels of first-generation admixture, indicating that a large proportion of the study group was made up of children of interracial families. I unfortunately didn’t have time to discuss the issue with them in depth, so I can only guess about the underlying cause of the challenges that they encountered. One of the fundamental assumptions for the statistical tests used in GWAS is that the subjects come from an ethnically homogeneous, random-mating population. When the population is not homogeneous, it is common to use a principal components correction to adjust for ethnic stratification. When we examine an individual admixed genome, like Blaine’s, we begin to see the flaws in that technique. Suppose we were conducting a GWAS in a primarily European population. Under the common approach of using genome-wide principal components to correct the test results, Blaine’s contribution to a test in regions where his genome is primarily European would be over-corrected, while the tests in regions where he has non-European DNA would be under-corrected. This issue may not cause any obvious problems if the total sample size is very large and the admixture is limited, but as sample size decreases and the extent of the admixture increases, the likelihood also increases that the test results will be confounded, despite (or perhaps even as a result of) PCA correction. In my colleagues’ project, the subjects with first-generation admixture had the entire length of their maternal and paternal haplotypes coming from different source populations, and the genome-wide correction factor may not have been ideal at any locus. The best way to analyze this data might be to separate the maternal and paternal chromosomes and calculate the correction factors separately for each haploid genome. Such an analysis approach would be difficult at best.
The best way to prevent analysis difficulties related to admixture is through careful planning and study design, ensuring that your samples are drawn from a homogeneous population. Unfortunately, that is not always possible. Most people don’t have the knowledge of their family history that Blaine has, making it difficult to collect and avoid unexpected admixture. Further, social behaviors in America and elsewhere have evolved such in recent decades that first-generation admixture is becoming more and more common. Admixture is something that will be ever more difficult to avoid in human subjects research, and it is something that we all need to understand better if we are to unravel the secrets of the genome. Much of the work we do in sequence analysis now revolves around characterizing rare variants, and familiarity with the subject’s genealogy is important to understanding the consequences of those variants and determining if they are even “rare” at all.
I would like to thank Blaine Bettinger for letting us all have a peek at his genome, and I wish him the best in his family history research. I’ve never been genotyped, but I would be surprised if my Y haplogroup was anything other than Danish. There are a few amateur genealogists in my family, and I’ve seen how much work goes into finding records for long lost ancestors and understand the tremendous satisfaction that comes with success. Perhaps someday genomics will help us all to better understand our history and how we are all related.
…And that’s my 2 SNPs.
There is definately a great deal to learn about this issue.
I really like all the points you made.