As VarSeq has been evaluated and chosen by more and more clinical labs, I have come to respect how unique each lab’s analytical use cases are.
Different labs may specialize in cancer therapy management, specific hereditary disorders, focused gene panels or whole exomes. Some may expect to spend just minutes validating the analytics and the presence or absence of well-characterized variants. Others expect every case to have its unique aspects, a puzzle to unravel with as many resources as possible as guides.
Yet for all the differences, these analytic workflows share quite a few key requirements:
- Detailed logs of the actions the user performed, allowing a trail of providence for the data evidence supporting their decision-making process
- A way for the hard work of variant assessments to be globally saved and applied to all newly imported variants
- Note taking on the context and insights surrounding a patient, with annotations being pulled in for a variant, its visual evidence and genomic context and snippets of web resources
- Ways of reproducibly filtering and ranking variants by bioinformatics thresholds, public annotations and known phenotypic information about patients
Filter, Sort, Rinse, Repeat
One thing you can expect when looking at NGS variant data is that no two tests will have the same bioinformatics. The differences may come from the sequencing machine, the chosen alignment and variant calling methods or the selection of target capture and read depth profile. Each test requires some tuning of your parameters to validate your QC and functional filters of variants.
In some cases where many genes are included (such as large gene panels, exome or genome sequencing), it’s useful to rank your high-quality variants against known phenotypic attributes about your samples. This is especially helpful when looking at trios or nuclear families with undiagnosed hereditary diseases.
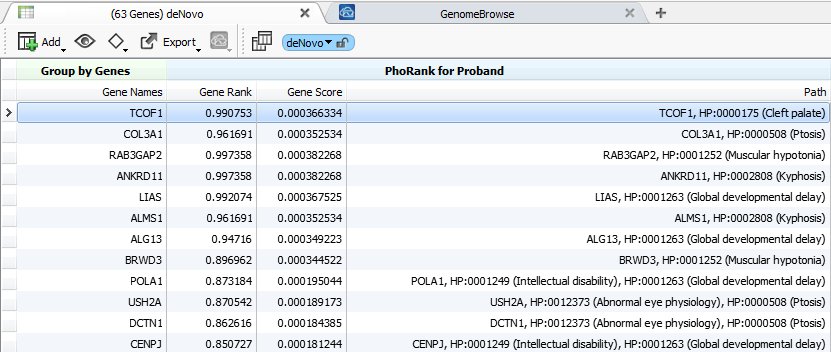
With the recent introduction of the PhoRank algorithm in VarSeq, we can leverage this phenotype information to integrate filtering and ranking approaches. Even as supplementary information to your current workflows, it’s informative to have a metric for the strength of known clinical associations between a gene you are investigating and the phenotypes available for a patient.
Once you have validated the filtering criteria for a new test, every annotation and choice you made is captured when you save your project as a Project Template, ready for new batches of samples to repeat the workflow as new projects.
And now, with the new Logs tab available at any time, you can drill into the history of each user action and analytical step taken on your projects.
Digital Lab Notebooks and One-Click Lab Databases
Listening to our stakeholders and early users, we added two new features specifically to support clinical workflows in the latest VarSeq releases: Project Notes and Variant Assessment Databases.

Quite simply, variant interpretation is a process that results in some decisions being made about the validity and suspected functional impact of a variant in a given sample and their presenting phenotypes. We built a very flexible database integration right into VarSeq that allows you to create simple flat-file or server-based databases to store and retrieve these decisions and notes.
Appearing as any other annotation source, these database can be integrated into your standard annotation and filtering workflow to ensure you don’t miss previously assessed variants in your future projects.

While a variant database is a great place to store your categorical assessments, sign-off status and other things you may want to capture about a variant, the new project-local Notes view allows you to have a space for creating per-sample aggregates reports and grab pretty much anything you can see into its editor.
You can save your notes as PDF or HTML files, and create as many as you like. One of my favorite features is the ability to drop bookmarks into specific GenomeBrowse views as well as context-rich rendered images from the current GenomeBrowse view.
See the Webcast for More
If these workflows and ways in which VarSeq is able to flexibly build them are relevant to your work or you just want to see what this looks like in a live demonstration, register for my upcoming webinar VarSeq as a Clinical NGS Platform.
I hope to see you there!