I was definitely an early adopter when it comes to personal genomics. In a recent email to their customer base announcing their one millionth customer, they revealed that I was customer #44,299.
And I have been consistently impressed with the product 23andMe provides through their web interface to make your hundreds of thousands of genotyped SNPs accessible and useful.
It takes an impressive amount of data science and statistical genetics to provide the trait and risk profiles they do. They also clearly take an engaged and caring stance on the ethics and risks of providing this data to users.
But one thing I’ve always really appreciated is their export function. From the “Browse Raw Data” view, there is a “Download” link that simply dumps those hundreds of thousands of genotypes into a simple plain text file.
That’s it. A file with your personal genotypes.
But what can you do with it?
Well… Now, you can use any of Golden Helix’s powerful products to open one or more 23andMe exports, and you can do quite a lot!
Personal Genomics 101
While we built VarSeq to provide a powerful mechanism to design and run reproducible variant analysis workflows for researchers and clinical labs, we have also focused on making it initiative and highly visual for those familiar with next generation sequencing, as well as for those who are not.
Golden Helix has put together a special program to support the education of the next generation of genomicists, and we have been thrilled to support Dr. Moore in using VarSeq to teach incoming freshman at UIUC genomics through a hands on and personal approach.

Figure 1: A student’s notebook capturing the investigation of a potentially interesting variant in VarSeq. It’s safe!
As part of the course, students get their own personal genotypes from 23andMe. With the latest version of VarSeq, they will be able to import this data directly to start doing the learning exercises that culminate in a personal case study.
Personal Data, Population Genetics
Along with my own data, I also have 23andMe data for my wife and first son. While I can hardly do all the great population genetics SVS has to offer, there are some methods that take a whole genome approach and work with common SNPs.
For example, along with various sample level QC, inheritance and Mendelian error checking, I can look for Runs of Homozygosity. These long stretches of homozygous genotypes indicate regions that may have copy number loss and should be inherited in a Mendelian fashion.
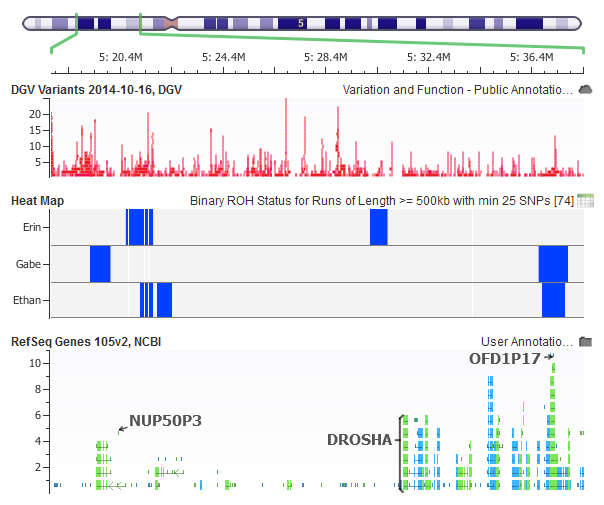
Figure 2: Inheritance of homozygous regions computed using SVS on my 23andMe genotypes. A region inherited from mom (top) and one from dad (middle) can be seen in the son (bottom).
Take a Peek, Get the Big Picture
While 23andMe data has huge value for educational purpose, there are a whole lot of citizen scientists and genomic enthusiasts that just like to dig into their own genomes.
Our popular free GenomeBrowse tool is commonly used by this crowd (along with researchers), and they have already requested an easier way to visualize 23andMe data in the context of all the rich public annotations GenomeBrowse has to offer.
With our upcoming GenomeBrowse release, they will have built-in support for converting the 23andMe text file into a plottable “Variant” source, capable of being searched, browsed and investigated.
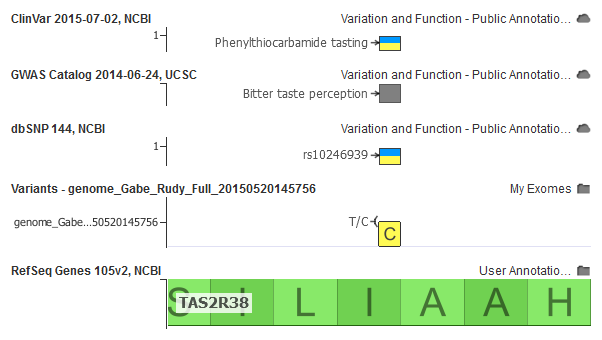
Figure 3: A GWAS study associates this variant in my 23andMe data with bitter taste perception.



1 Comments