GWAS became possible about 10 years ago as the result of several scientific advances. Since then, GWAS has continually developed as a primary method for identification of disease susceptibility genes in humans and other organisms.
At Golden Helix we are proud of our history in supporting GWAS analysis from its inception. Our software was used to analyze whole-genome data from the groundbreaking Affymetrix 10k SNP chip even before the chip’s commercial launch, and well before the NHGRI–EBI GWAS catalog keeps records. Among papers included in the GWAS catalog, Golden Helix citations appear as early as 2007.
I recently downloaded the GWAS catalog and assembled a few statistics that I would like to share. The GWAS catalog lists a 2005 Science report on age-related macular degeneration (AMD) as the first GWAS study, making 2015 the 10th anniversary of the modern GWAS. That study analyzed about 100k SNPs in just 146 study subjects. Despite a sample size that would generally be considered unacceptable today, the analysis successfully identified the CFH gene as a major AMD risk factor. The GWAS era was born as researchers rushed to replicate that success with other phenotypes.
The following table shows the number of GWAS studies published year-by-year from 2005 to 2013, based on counting unique PubMed IDs in the catalog. It also shows the mean and median sample size for the studies. The table only includes numbers for the primary analysis cohort – replication samples are not included.
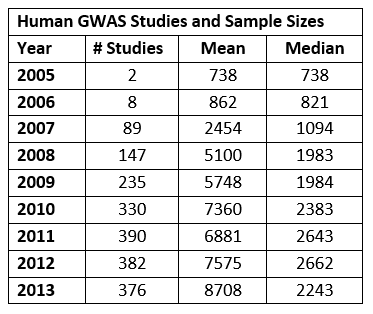
The table suggests that the volume of published human GWAS studies has plateaued in recent years, but the average size of the study cohorts continues to grow. The larger sample sizes can be attributed to many factors, including lower genotyping costs and researchers forming consortia to pool data and improve statistical power.
The figure below shows the growth in raw sample numbers, based on simply adding up the reported sample sizes for every study. About 70% of GWAS publications in the catalog also include a replication sample in addition to the main discovery cohort. The total number of samples used in replication sets is also shown.
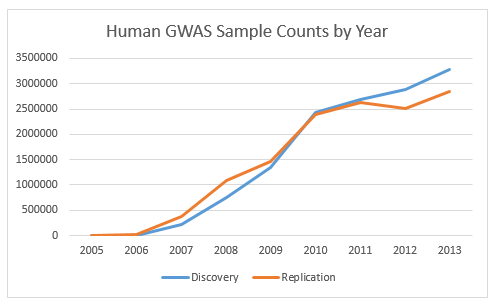
It is clear from the data that human genotyping arrays are being used in large numbers. About 60 million samples were included in GWAS studies in 2013. (Many samples are re-used in multiple studies, often as controls or replication samples, so the data should be interpreted accordingly.)
More recently, GWAS technology is experiencing growth in non-human applications, particularly in the burgeoning field of agrigenomics. High-throughput genotyping arrays are now available for numerous animals and crop species, enabling scientists and farmers to improve breeding programs and food production through genetics.
I’m not aware of an equivalent to the GWAS catalog for non-human applications, but it is informative to review the number of results returned by searching various terms on Google Scholar. I selected a few animal and plant species for which GWAS chips are available, and performed year-by-year searches to show the trends, as seen in the following table:

The numbers don’t all represent individual GWAS reports. They also include some review articles, methodology papers and conference abstracts. But the trend is clear: GWAS is growing rapidly in agricultural applications and shows no sign of stopping.
With 10 years of GWAS in the rear-view mirror, I think that the future looks bright. Many questions remain unanswered in genomics, and GWAS will continue to be an effective method to address those questions. At Golden Helix, we look forward to whatever challenges the future may have in store. Please contact us if you would like to discuss how we might be able to help each other.