
Short-read sequencing often fails to capture clinically actionable information from challenging genomic regions. While long-read sequencing now enables accurate and comprehensive detection of complex variants, there are still regions of the genome that remain notoriously difficult to analyze. Unfortunately, many of these regions contain genes that are highly relevant to precision medicine.
Consider CYP2D6 and CYP2C19, which guide pharmacogenomic dosing decisions; the HLA gene family, critical for transplant compatibility; PMS2 in Lynch syndrome; and SMN1 in spinal muscular atrophy, where pseudogenes create diagnostic confusion. These regions contain highly similar sequences arising from segmental duplications and gene family evolution, including gene birth, gene loss, and pseudogenes, which confound standard alignment algorithms and lead to missed variants and incomplete genotyping.
Developed by PacBio, Paraphase takes HiFi-aligned BAM files as input and addresses regions where variant calling is limited by high sequence homology. Paraphase takes a gene-family-centered approach, collecting all reads from a gene family and realigning them to a representative gene. Next, reads are phased into haplotypes, allowing Paraphase to examine all copies within a family, account for copy-number differences, and generate phased variant calls. This is critical for accurately analyzing many medically relevant genes, so we’ve incorporated it into automated workflows in VSWarehouse 3.
Integration in VSWarehouse 3
You don’t need to be a bioinformatics expert to benefit from powerful tools like Paraphase. VSWarehouse brings bioinformatics workflows directly to you in a user-friendly interface. Paraphase runs automatically as part of the HiFi Whole Genome Sequencing pipeline and PureTarget Carrier Panels workflow, which is available through the Workflow Registry and includes all prerequisites already accounted for in your VSW3 deployment. Select your input folder containing the data, an output folder, and hit Run.
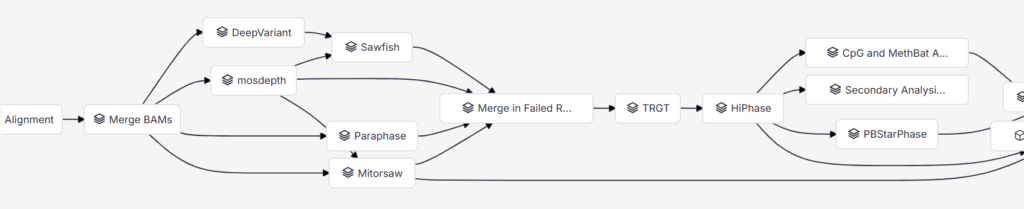
Tertiary analysis integration is simple with VSPipeline attached at the end of the workflow so you can incorporate Paraphase results into ready-to-interpret projects and draft clinical reports.
This automation eliminates the manual step of importing into VarSeq, providing a direct path from sequencer output to clinical interpretation. Paraphase integration supports both whole-genome sequencing and targeted sequencing data from PureTarget panels.
Conclusion
By integrating Paraphase into the VSWarehouse 3 PacBio workflows, Golden Helix enables more complete and accurate variant detection and genotyping to be automatically linked to tertiary analysis, reducing the diagnostic blind spots that arise from the idiosyncrasies of gene duplication and deletion and tandem gene expansions.