Golden Helix offers a market-leading bioinformatics solution that allows users to evaluate next-generation sequencing variants according to the ACMG and now ACGS guidelines. The ACMG guidelines were created in 2015 and are widely accepted as best practice for the interpretation of sequencing variants throughout the United States (Richard et al 2015). Very similar are the ACGS guidelines that were developed in 2020, which are primarily implemented in the European Union (Ellard et al 2020).
In 2018, the ACMG guidelines were tested under a Bayesian framework, which identified subtle inconsistencies in the rule logic for pathogenic and likely pathogenic variants (Tavtigian et al 2018). Specifically, this study focused on the combining of criteria for ACMG guidelines likely pathogenic rule (i) and pathogenic rule (ii), Figure 1. Using the Bayesian framework, likely pathogenic rule (i) had a higher increased posterior probability than pathogenic rule (ii) (Tavtigian et al 2018). This indicated that likely pathogenic rule (i) had a higher level of pathogenicity than pathogenic rule (ii) of the ACMG guidelines. This was also supported by the fact that variants that have one very strong and one moderate evidence of pathogenicity (likely pathogenic rule (i)) tend to be loss of function variants that lead to nonsense-mediated decay of the transcript (Ellard et al 2020).

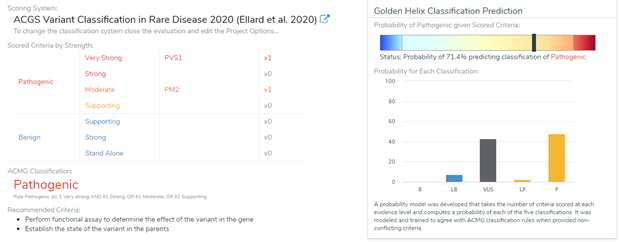
Using these results, the Association for Clinical Genomic Science (ACGS) made updates to the ACMG recommendations. As shown in Figure 2, one very strong and one moderate level of evidence is classified as pathogenic, whereas only two strong evidence of pathogenicity results in a likely pathogenic classification (Ellard et al 2020).

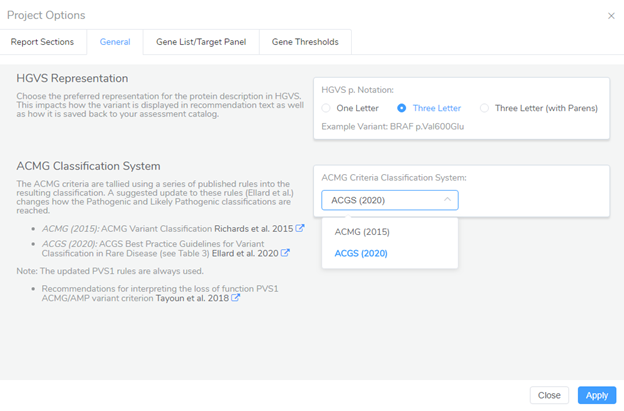
With the new release of VarSeq, updates to PVS1 are now incorporated into the ACMG guidelines of VSClinical and users can also use the ACGS guidelines for variant evaluation. The option to select between the ACMG and ACGS guidelines is in the Project Options of VSClinical and should be selected prior to starting an evaluation, Figure 3. Once the classification system is selected, it will be applied in the VSClinical interpretation hub and will apply the scoring criteria for these guidelines, Figure 4.
Together, VarSeq v2.2.2 incorporates many new features including the ability to use the ACMG or ACGS guidelines for the interpretation of sequencing variants. We will be discussing other new features provided in this release but if you have any questions about the updates or the software in general, you can reach out to [email protected]. Feel free to check out some of our other blogs that always contain important news and updates for the next-gen sequencing community.
Works Cited:
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL; ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015 May;17(5):405-24. doi: 10.1038/gim.2015.30. Epub 2015 Mar 5. PubMed PMID: 25741868.
Tavtigian SV, Greenblatt MS, Harrison SM, Nussbaum RL, Prabhu SA, Boucher KM, Biesecker LG; ClinGen Sequence Variant Interpretation Working Group (ClinGen SVI). Modeling the ACMG/AMP variant classification guidelines as a Bayesian classification framework. Genet Med. 2018 Sep;20(9):1054-1060. doi:10.1038/gim.2017.210. Epub 2018 Jan 4. PubMed PMID: 29300386.
Ellard S, Emma B, Callaway A, Berry I, Forrester N, Turnbull C, Owens M, Eccles D, Abbs S, Scott R, Deans Z, Lester T, Campbel J, Newman W, Ramsden S, McMullan D. ACGS Best Practice Guidelines for Variant Classification in Rare Disease 2020. Association for Clinical Genomic Science.


