Bioinformatics Pipelines and Systems Infrastructure
The genetics industry is undergoing a fundamental shift from a clinical science focus to a bioinformatics focus. Genetic services require a greater level of data analytics sophistication than is required for other laboratory testing. Currently, data generated by new tests overwhelms current information technology systems and human interpretation capabilities. This is one of the reasons that we at Golden Helix strive to simplify the process of analyzing and interpreting the data, so that it is possible for a wider group of users to conduct work in this space.
Ultimately, the output of the NGS data analysis pipeline needs to be integrated into the electronic health record and to be aggregated across a patient population. Robust informatics systems and trained bioinformaticians are critical new additions to the clinical team, underscoring the growing importance of clinical bioinformatics infrastructure. Servant et al. (2014) covered this issue in detail. I agree with their findings. Here is the upshot.
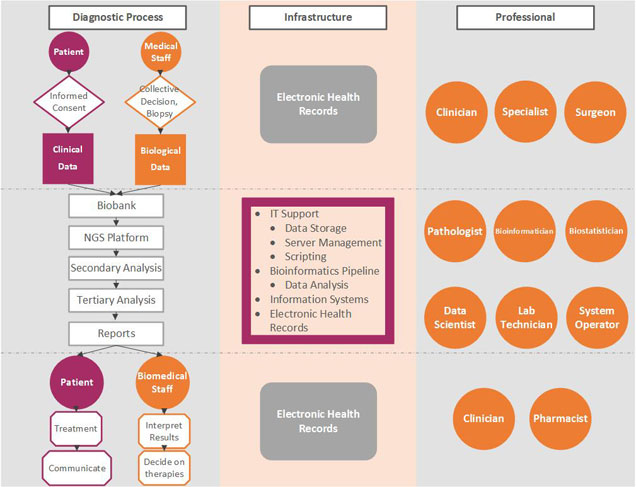
Precision medicine requires a strong interdisciplinary collaboration between several stakeholders covering a large continuum of expertise ranging from medical, clinical, biological, translational, technical, and biotechnological know-hows. The chart below illustrates the different practitioners involved in the complex process, describes the data workflow starting from and coming back to the patient in order to tailor the therapy and shows the informatics and bioinformatics infrastructure supporting the workflow. To build the therapeutic decision, the most exhaustive data ranging from clinical to biological, environmental and family information (e.g., description of the tumor histology, list of previous treatments, family history, etc.) needs to be collected along a complex healthcare pathway. During the process, physicians (such as surgeons, pathologists, radiation and medical oncologists, etc.), biologists, pharmacists, bioinformaticians, computational biologists, biostatisticians, biobank managers, biotechnological platform managers, clinical research associates, and the technical staff will offer their expertise for the benefit of the patient.
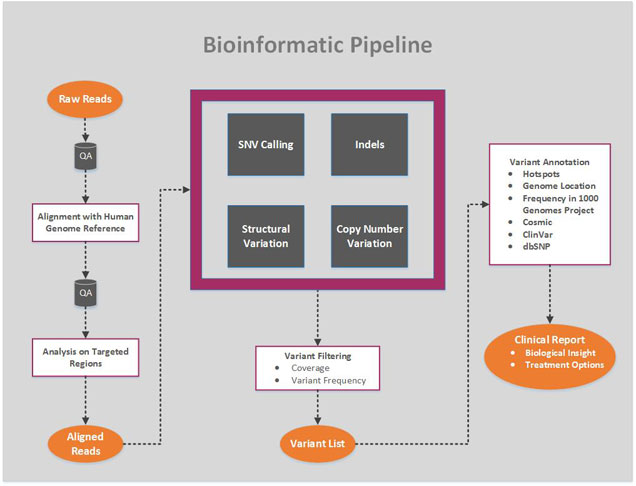
The second figure outlines the general structure of a bioinformatics pipeline designed to detect somatic mutations — a detailed example of somatic analysis — as an illustration of the building blocks of a bioinformatics pipeline. The reads could come from a number of different sequencers such as the Ion TorrentTM PGM sequencer using the AmpliseqTM cancer panel. The reads are aligned on the reference human genome. Then, in the next step variants have to be called (SNVs, indels and possible more complex variations like structural variations and copy number alterations). Variants have to be filtered to promote a high specificity, in order to avoid any false positives. Thus, detected variants are filtered according to their frequency (e.g ≥4% for SNVs and 5% for indels, strand ratio ≥0.2, and reads coverage ≥30X for SNVs and 100X for indels). In addition, SNVs and mainly insertions and deletions detected in the context of a repeated region or a homopolymer are double checked. Homopolymer and repeated regions are prone to contain recurrent false positive, because of the limitation of the current sequencing technology. To facilitate the interpretation of individual patient data for clinical trials, the filtered list of variants is then annotated using software packages like Golden Helix’ VarSeq for variant annotation. Common polymorphisms as well as recurrent and neutral variants on hotspots are reported. These variants do not present any therapeutic interest but are good internal controls to ensure the quality of the sequencing data. The Catalog of Somatic Mutation in Cancer (COSMIC) is used to annotate the mutations detected at a hotspot position. Non targeted mutations in genes covered by the panel, being non polymorphic nonsense, missense or indels are also reported, even if it may be difficult to know whether the alteration is involved in deregulating a particular pathway and whether it is clinically relevant. However, more stringent frequency filtering are applied for these cases (e.g. frequency ≥10% for SNVs and 15% for indels) leading to a higher specificity. Then, relevant mutations and variations are visualized using the GoldenHelix’ GenomeBrowse. The visualization remains an important step to assess the overall quality of the variant call, by taking into account the reads coverage, the error rate in the flanking region, the mutation position across the targeted region and across reads supporting them.
The evolution of sequencing technologies rapidly expanded the frontiers of genomics into clinic at lower costs and increased speeds. Bioinformatics pipeline will become part of the standard process to diagnose patients and to determine treatment options. Our understanding of how this looks like in the oncology space — supported by dedicated oncology software — is already reasonably developed. Going forward we will have to standardize these pipelines and embed their output into patient heath record systems.