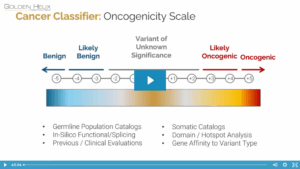
The first two blogs in this series covered Sentieon’s somatic variant calling from Tumor with Normal (Part I) and Tumor without Normal (Part II). In addition to providing multiple somatic variant calling processes, users also have access to high-sensitivity scripts and full support for the GRCh38 reference assembly. Without going into excessive detail, this final blog of the series will cover the highlights of these additional features available to customize any NGS-based somatic analysis workflow.
The pursuit of somatic variant detection is tricky for many reasons. For example, the nature of having ample coverage deep enough to determine abnormal mutation allele frequency. From a secondary analysis perspective, the process of alignment and variant calling undergoes many quality control steps. Ultimately, most variants listed in the VCF likely have a decent probability of being real and not simply artifacts. However, it is the case some questionable variants may be real and a clinician may wish to reduce the strictness of these quality controls steps and probabilities to capture any possible somatic variant. Luckily, with Sentieon, users have full control of these algorithms to increase sensitivity. Moreover, the nature of processing the increased number of variants in the VCF can be handled effectively in VarSeq as a tertiary analysis solution. Golden Helix has developed example high-sensitivity scripts that we can provide to those testing or using Sentieon for somatic calling as a starting guide to further customize their desired detection level.
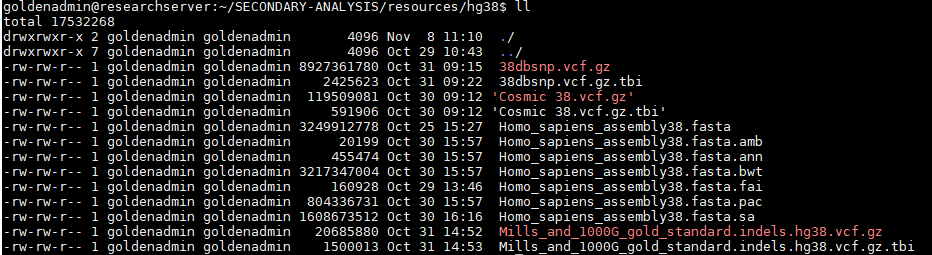
It may also be the case that users wish to generate their NGS data based on the most recent GRCh38 reference assembly. Golden Helix now hosts the 38.fasta for alignment with various annotations (dbSNP and Cosmic), and known indels for indel realignment (Mills and 1000G gold standard).
With GHI solutions, users can deploy Sentieon’s top-quality somatic variant calling and subsequently perform tertiary analysis with our VSClinical AMP Workflow. If you are interested in setting up a full NGS bioinformatic processing architecture, please reach out to us and we will help you get started!