In a previous blog post, I demonstrated using VarSeq to directly analyze the whole genome sequencing data of 17 supercentenarians. Since then, I have been working with the variant set from these long-lived genomes to prepare a public data track useful for annotation and filtering.
Well, we just published the track last week, and I’m excited to share some of the details involved in its making.
The track, named Supercentenarian 17 Variant Frequencies, GHI, provides not only the allelic frequency of observed variants in these 17 whole genomes, but also the counts of the heterozygous and homozygous genotypes for those individuals. This kind of population-level data is invaluable for variant annotation and prioritization workflows.
For example, when investigating a rare recessive disease, its probably safe to say any variant occurring in a homozygous state in a 110 year old individual is probably not your causal disease mutation.
So what was tricky about constructing this population variant catalog?
It turns out, quite a lot.
Multi-Allelic Sites and Population Catalogs
When annotating variants from your own data against a population catalog, you want the catalog to have the most precise set of information for the set of alleles you observe in your data.
For example, if you see a heterozygous A/C (where A is the reference allele) at a given site, you would like to see the allele counts and frequencies of the “C” as well as how many times the A/C het or “C/C” homozygous variant occurs.
But if the population catalog contains some samples that have an A/C at that site, some that have an A/T and then one that has a C/T (two non-ref alleles), the variant caller will place all of these in a single A/C/T record in the VCF file.
When importing data into VarSeq using the Individual Samples mode in the import wizard, we by default will have selected the Advanced Option of Split Variants Based on Unique Genotypes (described in our manual here), that will split the A/C/T record into ones that succinctly represent the samples with the A/C, the A/T and the C/T forms of the variant.
In more complex examples, breaking out these multiple alternates into their own records can change the representation of variants to match how they would be independently called.
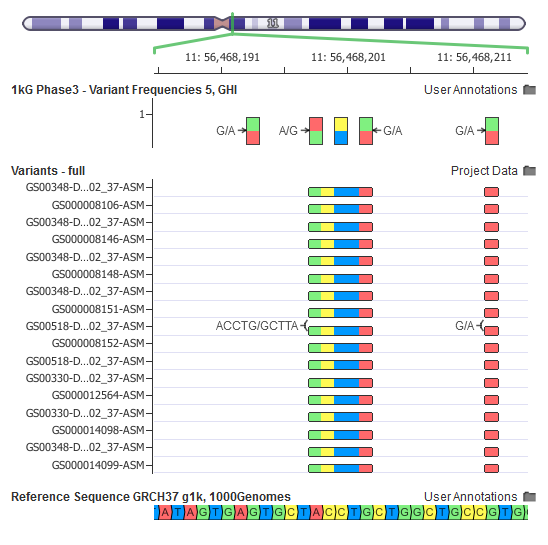
In this example, is the A/T variant in the bottom variant map present in the ExAC catalog?

In the raw VCF from the ExAC FTP site (middle track), a variant site contains two alternates: TTC, A. But once we do a multi-allelic split and normalize the variant representation (top track), we have two adjacent variants of A/T and TC/- (deletion of a TC).
With the ExAC catalog in this form that we use in our public repository, we can clearly annotate our A/T variant.
Allelic Primitives And Left Align: Normalizing Variants for Annotation
Two other advanced options in that final pane of the import wizard are:
- Allelic Primitives: Split multi-nucleotide polymorphisms (MNPs) into individual bases and insertions and deletions.
- Left Align: Shift variants to their left-most representation on the reference genome (using a Smith-Waterman local realignment)
These are both applicable to the Supercentenarian variants, due to the preference of the complete genomics variant caller to call mutations within two base-pairs of each other as a single mutation event (creating MNPs).
Here are two examples of these options in action:
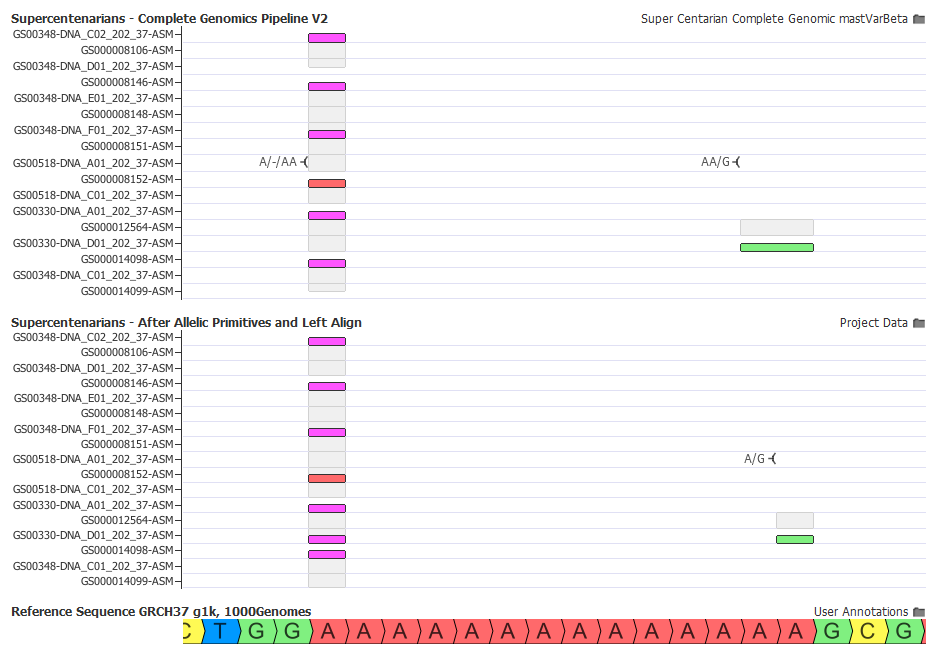
Ready for Annotation and Filtering
With the publication of this track, you can now visualize these variants directly from GenomeBrowse or use it for annotation and filtering in SVS and VarSeq — the kind of capability that sets purpose-built NGS analysis software apart.
In VarSeq, you will match variants containing the alleles present in samples, which will often but not always be a single record.
For example, at 6:29911119 in our tutorial trio project, there are two alternates of a C and a T, with the following genotype table:
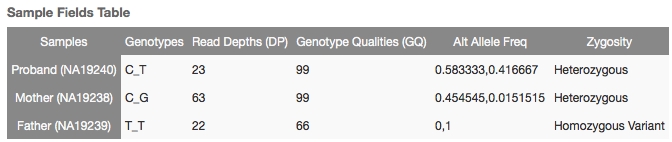
The Supercentenarian annotation found records for both alternates, and looks like this:
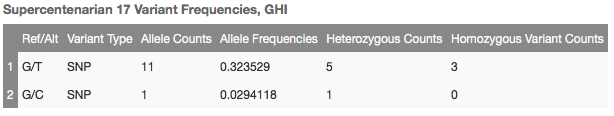
Whether filtering out likely benign variants or assisting in interpretation, this track is a valuable resource for tertiary analysis and contains some fascinating and useful information from a very select population.
The Secret to Long Life? It May Lurk in the DNA of the Oldest Among US – The New York Times https://www.nytimes.com/2017/11/13/health/supercentenarians-genetics-longevity.html