With the recent release of VarSeq 1.4.7, we have expanded the concepts of our popular assessment catalog to include CNV and other region-based records and not just variants.
To match these capabilities, we have made a major update to VSWarehouse that supports these new record types in the centrally hosted and versioned Catalogs and Reports.
Review of the VSWarehouse Genomic Data Store
VSWarehouse has three major resources that can be shared and managed on a single server: Projects, Reports, and Catalogs. Projects are the fully merged and annotated repository of every variant you sequence. This allows you to build up your own internal variant frequency annotations based on your laboratory samples, potentially broken down by different groups or cohorts.
Reports store the final output of the clinical interpretation process with per-sample background data, clinical outcome and the user-selected variants and CNVs to be included in our fully customizable rendered reports.
Catalogs provide a flexible knowledge base to capture variant and CNV interpretations, as well as lists of useful variants and CNVs to flag such as recurring false-positives, confirmed benign variants etc.
Store, Annotate and Plot your High-Quality CNVs in VSWarehouse
After doing some annotation and filtering of the CNVs called by the latest VarSeq VS-CNV algorithm, you will find that there are relatively few CNVs occur for each gene panel or exome-based sample. For this reason, you may want to catalog every high-quality CNV observed in your clinical test.
With the new VSWarehouse sample-keyed catalog and ability to annotate against CNV based assessment catalog, you can now build up your own “Observed CNV” annotation source very similar population catalogs such as the ExAC XHMM CNV Calls.
To set up your own Observed CNV catalog, we first go to the VSWarehouse manager and click “Create” under the Catalogs tab. Select “CNV” as the catalog type, and also set a Custom Key that captures the current Samples name:
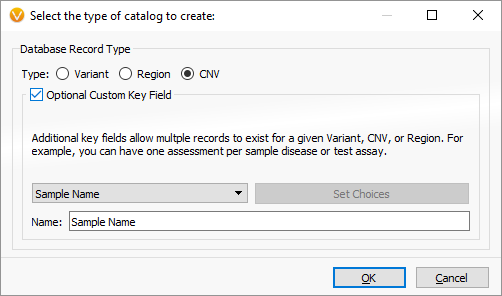
By using the Custom Key field, we can store multiple records in the catalog that share the same genomic coordinates as long as they are detected in different samples. This allows us to build up the catalog and use it as a “CNV frequency” annotation source.
In the next window, we can use the pre-built schema “Sample CNV State” to pre-fill in our schema with fields we can capture from the CNV table. Feel free to customize these schema fields, but also realize you can change them at any time in the future!
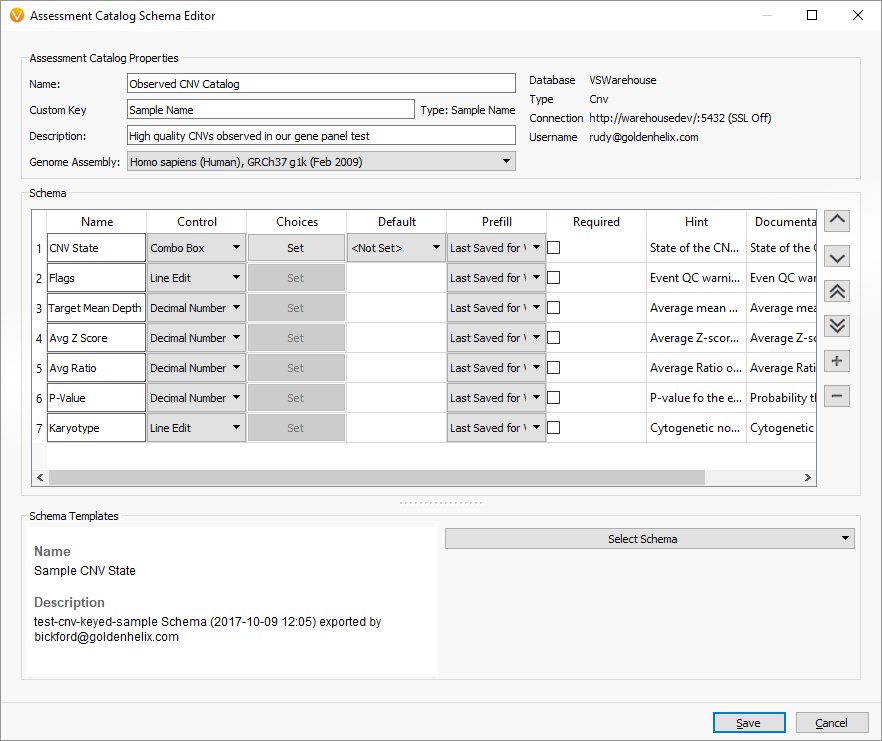
After creating this catalog, select “Open” to start using it as you would any Assessment Catalog when selecting CNVs to interpret.
To upload all the filtered variants in your CNV table for the current sample, simply go to the new Import -> Import from table menu on the opened Catalog tab and follow the wizard. This is the equivalent of selecting each CNV in your table and interactively adding its data to the catalog.
After doing this on a few samples, your VSWarehouse catalog will start to look something like this:
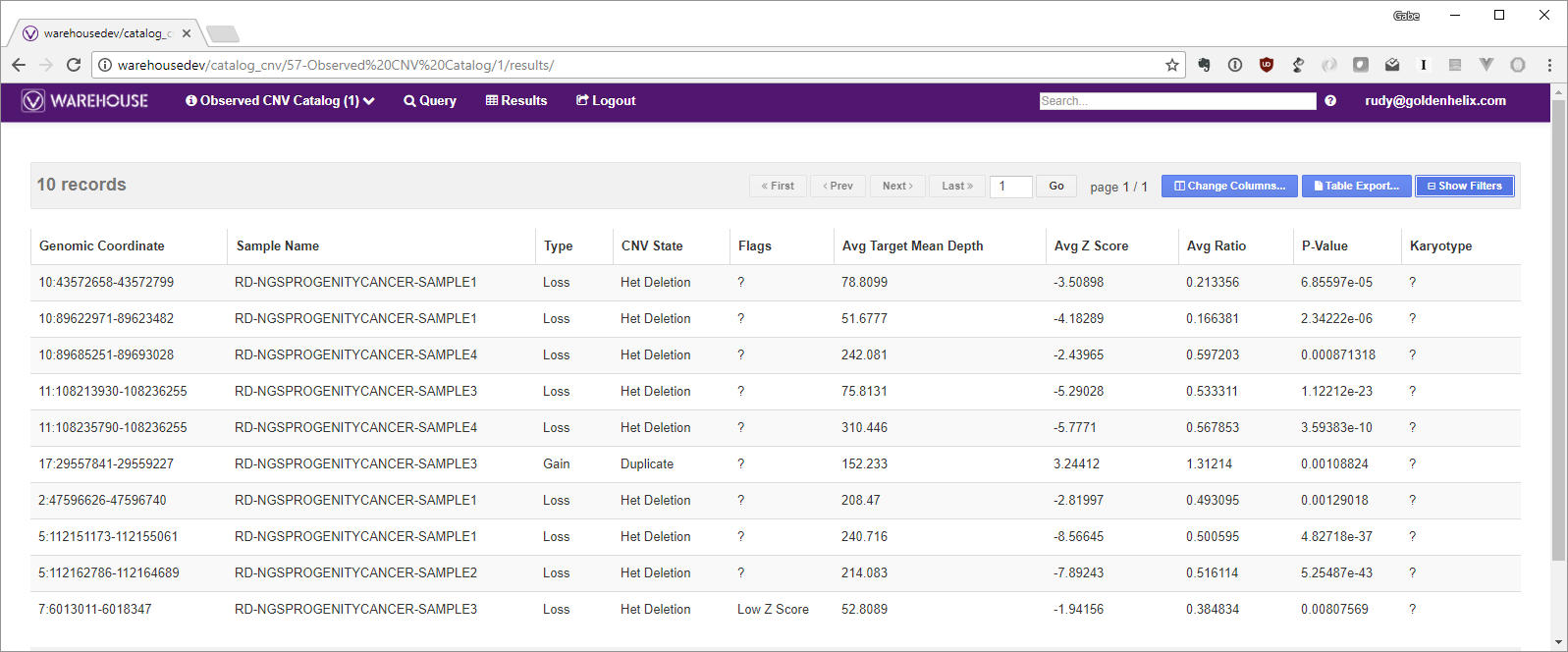
The next step is to add this catalog as an annotation source to your CNV table so you can start to see when CNVs in new samples overlap or match those in your Observed CNV Catalog!
From the top-level Add toolbar menu, go to Secondary Tables -> CNV Annotations and select VSWarehouse server as the Location and the new Observed CNV Catalog as the annotation source.
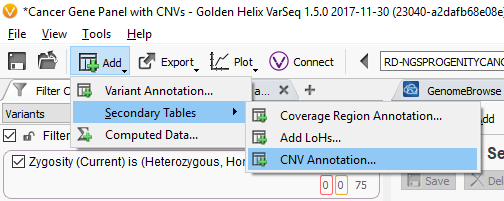
Similarly, you can add your Observed CNV Catalog to your GenomeBrowse view and plot all the records from your catalog along with the evidence and annotations of your current sample.
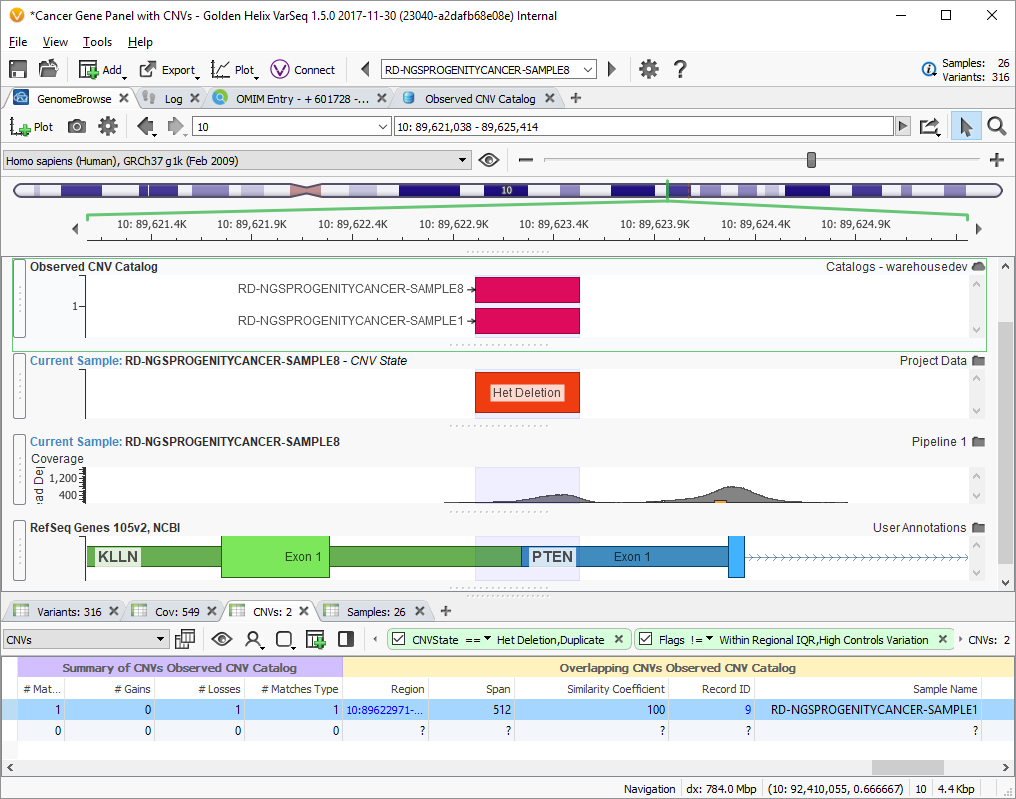
For example, in this screenshot, we can see a Het Deletion in PTEN that is also in another sample in our catalog. This shows up in the CNV table since we added it as an annotation source as well. You can see in the Summary columns that we have one matching CNV. In the second set of columns, we can find per-sample details and drill down links into VSWarehouse to learn more about the CNV calls and the sample they were observed in.
Warehouse as a Source of Truth
When designing complex data systems, it is considered best practice to have a single “source of truth” from which secondary systems may operate on, creating their own views or derived analytics. The same concept applies to the data accumulated during the analysis of a clinical genomic test.
With this update, we are responding to our users who want to keep regional and CNV-based genomic annotations and clinical interpretations in VSWarehouse as the single source of truth.
If you are looking for a solution to be the single source of truth for your genomic laboratory data, scaling with your lab from dozens to tens of thousands of samples, please reach out and we will be happy to provide you with a demo and customized evaluation experience!


