
Short Tandem Repeats (STRs) are increasingly recognized as clinically important genetic variants, contributing to a wide range of rare inherited disorders. Historically, the analysis and clinical reporting of STRs have been challenging due to limitations in both sequencing technologies and interpretation frameworks. With the advent of long-read sequencing technologies and the advanced secondary analysis pipelines provided by PacBio, Oxford Nanopore, and DRAGEN, clinicians now have reliable methods to detect STRs. However, the process of interpreting these mutations remains challenging.
We are excited to announce that VarSeq now offers a streamlined workflow for analyzing and reporting STRs alongside small variants. By leveraging annotations provided by STRchive, VarSeq enables users to generate detailed and standardized clinical reports that combine both small variant and tandem repeat findings.
In this blog post, we will walk through how VarSeq can be used to identify clinically relevant STRs and generate a professional clinical report that incorporates both small variants and STR findings.
Annotating and Filtering STRs in VarSeq
For our example report, we will analyze a sample dataset that includes small variants and STR calls produced by a DRAGEN secondary analysis pipeline. Both data types are imported into VarSeq using the shipped DRAGEN STR Template for GRCh38. This template produces two variant tables, each corresponding to a parallel filter chain.
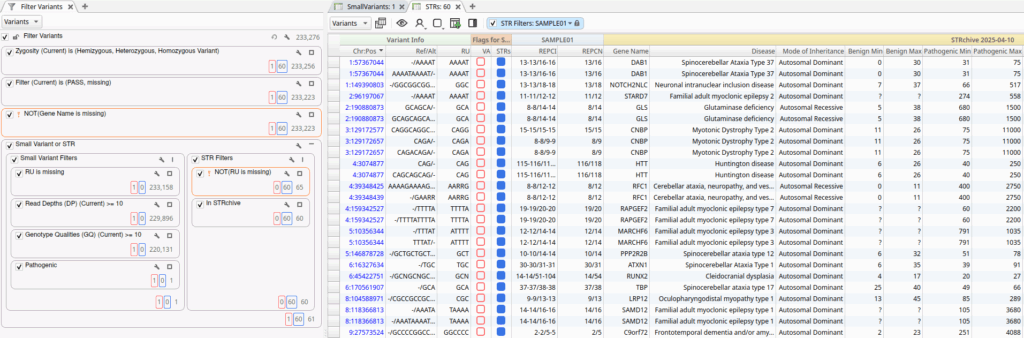
The small variant filter chain is designed to identify high-quality, clinically relevant small variants. It includes filters on basic quality fields, such as Read Depth and Genotype Quality, along with filters based on classifications provided ClinVar and VarSeq’s automated ACMG Classifier. In this example, the workflow identified a single Likely Pathogenic variant in the PPIP5K1P1-CATSPER2 gene.
The STR filter chain identifies variants with a valid repeat unit (RU) field that overlap repeat regions in the STRchive annotation source. For this dataset, the sample contained 60 STRs with pathogenicity information in STRchive. The filtered variants from both filter chains were added to the Variant and STR record sets, flagging them for inclusion in the clinical report.
Generating the Clinical Report
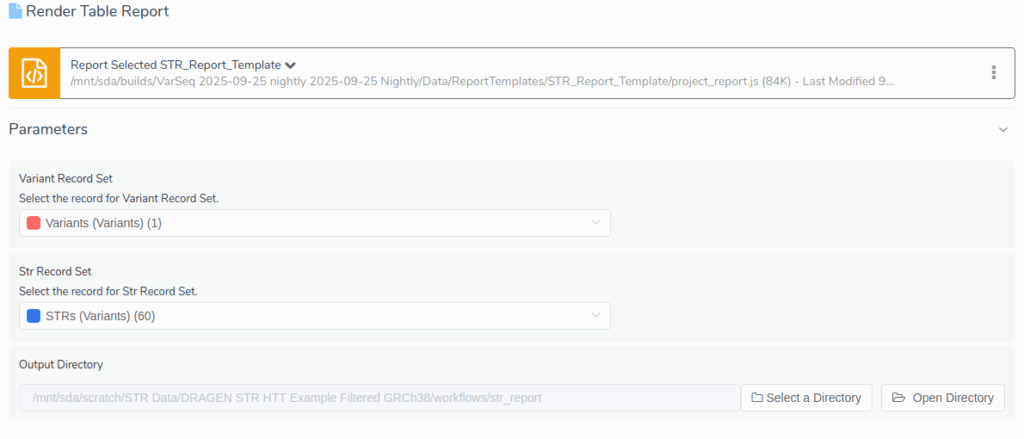
Once filtering and annotation are complete, VarSeq makes it simple to generate a clinical report. Using the Table Report interface, we selected the STR_Report_Template, which requires three input parameters:
- Variant Record Set: Specifies the set of small variants to report.
- STR Record Set: Specifies the set of STRs to report.
- Output Directory: Determines where the output files will be written (defaults to the project’s workflows directory).
Once we have specified the relevant parameters, we can generate the clinical report. At the top of the report, we can see variant summary information for the likely pathogenic variant that was flagged for inclusion, followed by the tandem repeat summary.
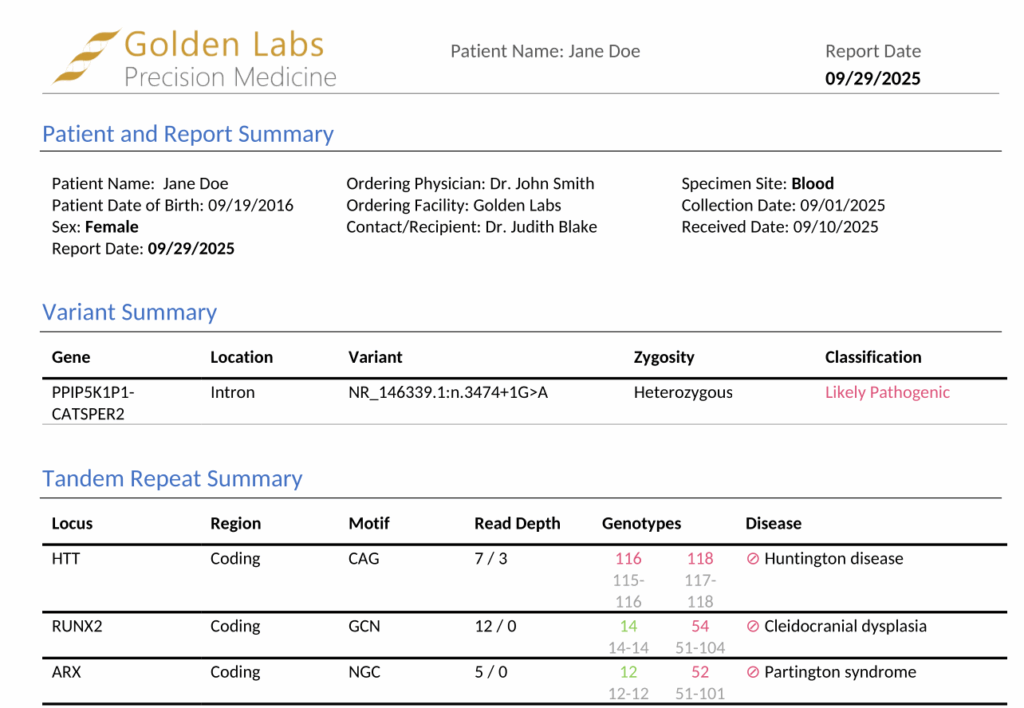
The tandem repeat summary presents critical information in an easily digestible format, including the associated gene, repeat motif, linked disease, and repeat counts for each reported STR. In our example, the summary reveals that this sample harbors homozygous pathogenic repeats in the HTT gene and is heterozygous for pathogenic repeats in both the RUNX2 and ARX genes.
Scrolling down, the Primary Findings section provides detailed information on each pathogenic STR. This includes essential clinical context such as Mode of Inheritance, Prevalence, and Age at Onset for the associated disease. These details are pulled directly from STRchive and include inline citations that link to relevant publications.
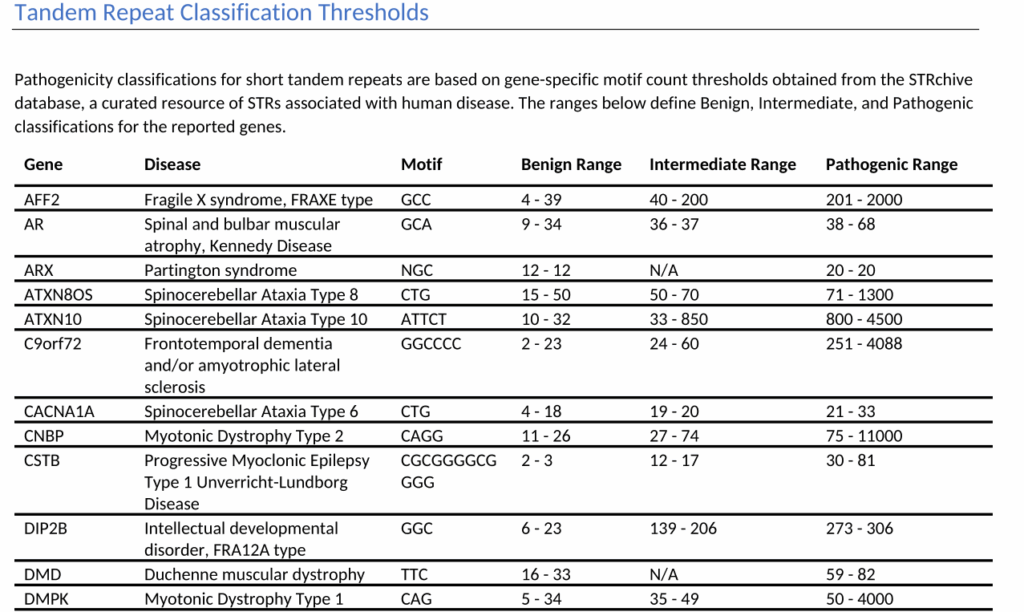
The report continues with a section documenting the tandem repeat count thresholds used to classify each reported STR. This transparency in classification criteria is essential for clinical interpretation and quality assurance. Following the thresholds, the report lists any incidental tandem repeats detected in the sample that were not associated with disease in STRchive.
Finally, the report concludes with a complete References section that contains all citations mentioned throughout the document, alongwith external links to PubMed for easy access to the primary literature.
Customization and Automation
While the shipped STR report template provides an excellent starting point, it’s important to note that both the underlying code that generates the report and the Microsoft Word template used to format the output are fully customizable. This flexibility allows laboratories to tailor the report format, content, and styling to meet their specific institutional requirements and branding preferences.
For laboratories processing high volumes of samples, the entire workflow described above can be automated entirely using VSPipeline, Golden Helix’s command-line automation tool. VSPipeline can be configured to automatically import variant data, apply the appropriate annotations, evaluate the filter chains to identify clinically relevant variants and STRs, and generate the final clinical report, all without manual intervention.
Conclusion
By combining the comprehensive STRchive annotation database with intuitive filtering workflows and automated report generation, VarSeq streamlines the process of STR interpretation and reporting. The software’s ability to generate reports with detailed clinical context, transparent classification criteria, and full citation tracking ensures that clinicians receive the information they need to make informed decisions. Please contact our team if you would like to know more about VarSeq’s STR reporting capabilities.
