Our tutorials are a popular resource for prospective users. We have added a tutorial to help introduce customers to the ease and utility of the ACMG Guidelines incorporated in VSClinical. The ACMG Guidelines are a collection of 33 criteria that are used to determine a variant’s level of pathogenicity.
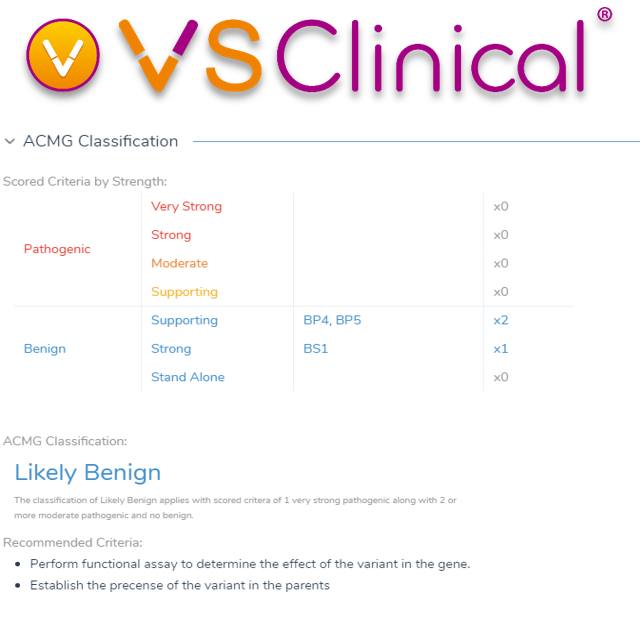
VSClinical and VarSeq make it easy for users to sort and filter variant sets, answer questions to analyze the ACMG criteria used to score and classify variants, and then store these assessments and classifications. This means that VSClinical allows for repeatable and streamlined workflows through this complex analysis from variant import to final report creation.
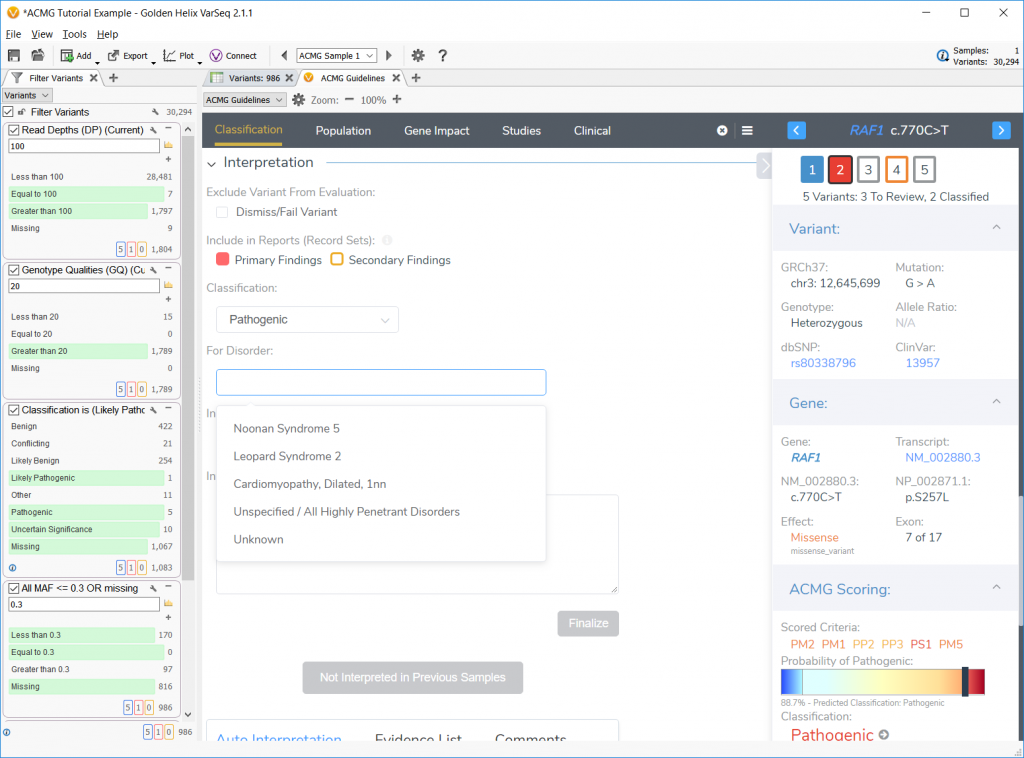
The new ACMG Guidelines tutorial will start off users with a blank project and guide them from the filtering of a provided sample VCF through the ACMG Guidelines analysis to a final report creation. The tutorial delves into 4 specific example cases. First, a benign frameshift in the MSH6 gene is analyzed and a straight-forward pathogenic classification of a missense variant in the RAF1 gene is presented.
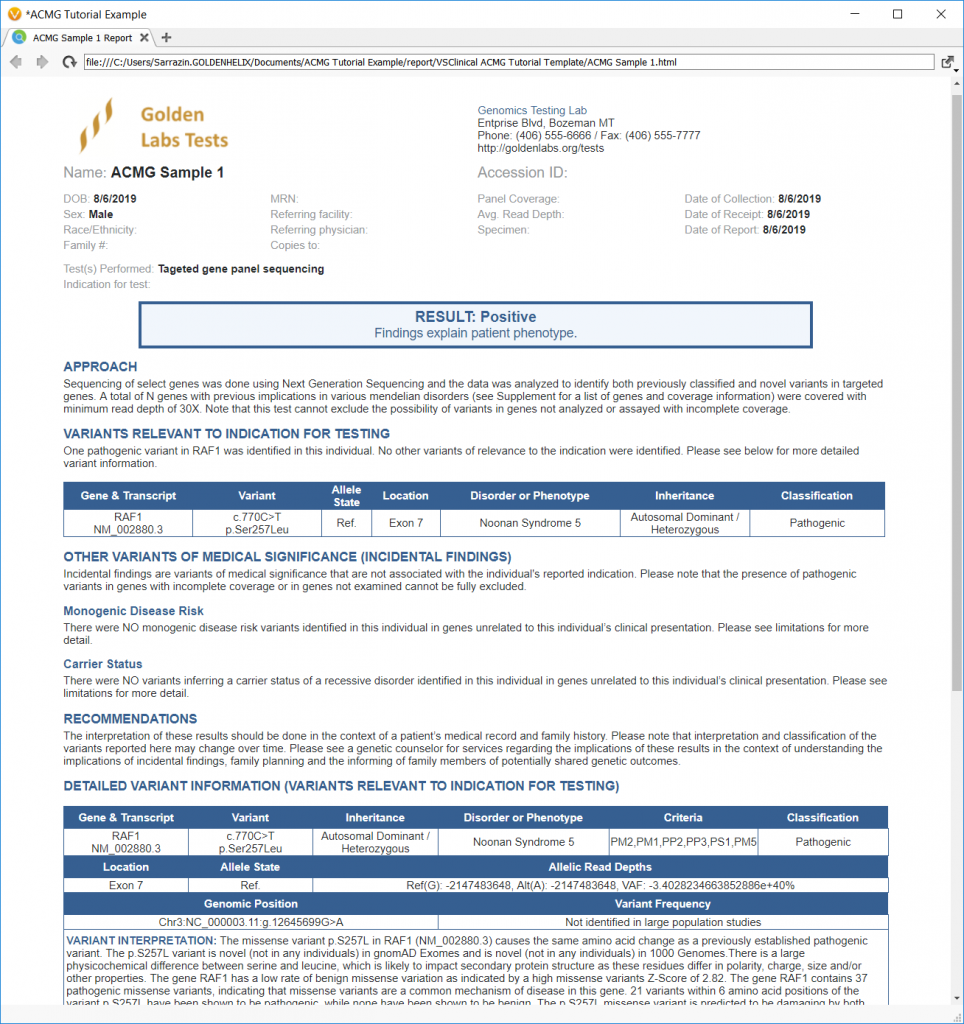
Then, a more edge-case PTEN stop gain and EGFR in-frame insertion are analyzed. Finally, the tutorial ends by showing how users can simply include different interpretation options and create a custom lab report.
We are excited to present this new tutorial for users to explore the features presented in VSClinical and incorporate the ACMG Guidelines in their variant analyses!
If you are interested in getting a demo license and jumping into the AMP Guidelines with VSClinical, please request a demo let us know here and we will help you get started.




Thank you for sharing this article. It is informative and new to me. Hope people like me will get to know more about this with the help of this article. Best wishes.