Golden Helix works to keep incorporating and updating great somatic annotation catalogs for our VSClinical users. We currently have the updated version of one of the largest cancer databases from the International Cancer Genome Consortium, or ICGC. Version 28 has been improved by integrating ClinVar and CIViC clinical annotations, and as always, increasing the number of mutations listed.
The current version (V28) consists of:
- Cancer projects: 86
- Cancer Primary sites: 22
- Donors with molecular data in DCC: 22,330
- Total Donors: 24,289
- Simple somatic mutations: 81,782,588
- Mutated genes: 57,905
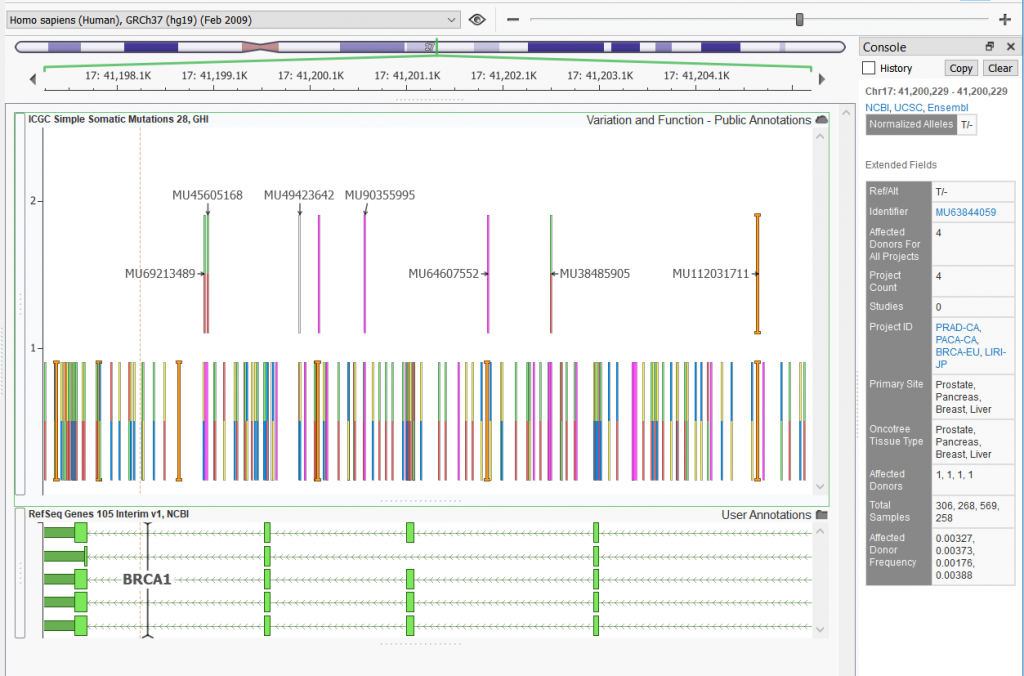
The image above shows part of the ICGC v28 track plotted in GenomeBrowse. ICGC continues to stand out as one of the largest catalogs of somatic mutations and is included in VarSeq’s standard list of annotations.
Another great cancer workflow annotation source that is part of VarSeq’s premium annotations is COSMIC. VarSeq currently hosts COSMIC v89 which pulled in data from over 8000 new samples compared to its previous release to make its sample count over 1.42 million and hosting over 6 million coding mutations. COSMIC includes more than just cataloged somatic mutations, and we curate a number of tracks based on the COSMIC databases, including:
- Cancer Gene Consensus: Properties and annotations on cancer genes
- Fusions: List of known translocations and gene fusions
- Hallmarks of Cancer: Descriptions and citations of how each cancer gene demonstrates the Hallmarks of Cancer and the functional pathways in which it is active
- Mutations: Mutation catalog derived from studies and whole-genome projects on tumors
- Transcript Counts: Categorizing on a transcript level the types of mutations and tumor sites of their origin.
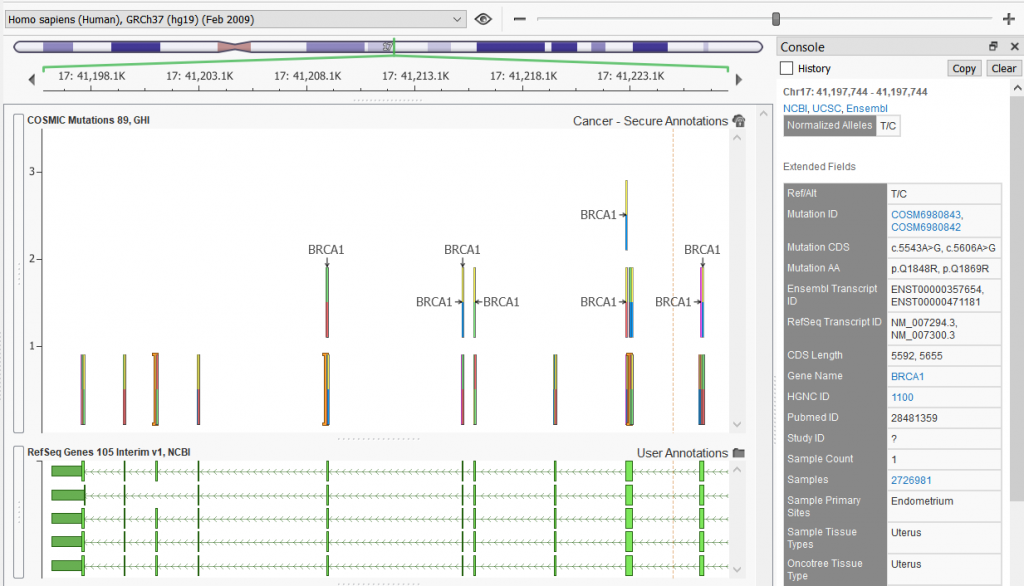
All of these tracks (as well as the ICGC track mentioned above) are available in both GRCh37 and GRCh38 genome assemblies. An example of the Mutations track plotted in GenomeBrowse is shown below.
Last month, COSMIC released version 90 and our curation team is already working on bringing this data to VarSeq users, so be on the lookout soon!
If you have any specific questions about access to VarSeq, demos, or cost, request more information here!