In this blog post, I’ll highlight the benefits of using VSPipeline to automate tertiary analysis in VSWarehouse. Users can now run VSPipeline project creation as a standalone task or a final step in a bioinformatic workflow without writing code. Simply use one of our shipped VarSeq projects or create your own, then automate that project as a template that can run on large batches of samples.
Breaking Through the Analysis Bottleneck
Modern sequencing technologies generate vast amounts of potentially useful data, but you might relate to the frustration that comes when FASTQ or VCF files begin to collect dust because of a bottleneck in your analysis procedures. The real crux isn’t producing the data – it’s skillfully moving the data into interpretation and reporting, so that accurate results can be communicated to patients and other stakeholders.
VSPipeline in VSWarehouse
VSPipeline is Golden Helix’s solution for automating tertiary analysis workflows. With VSPipeline, a VarSeq project can be used as a template for automatically carrying out variant import, annotation with clinical databases, variant prioritization through filters, and even interpretation of flagged variants and report generation. For years, VSPipeline has helped VarSeq users upgrade their diagnostic throughput as a command-line tool. Now, in VSWarehouse, VSPipeline has become even easier to use at scale, shipped as a VSWarehouse task.
What Are Tasks in VSWarehouse?
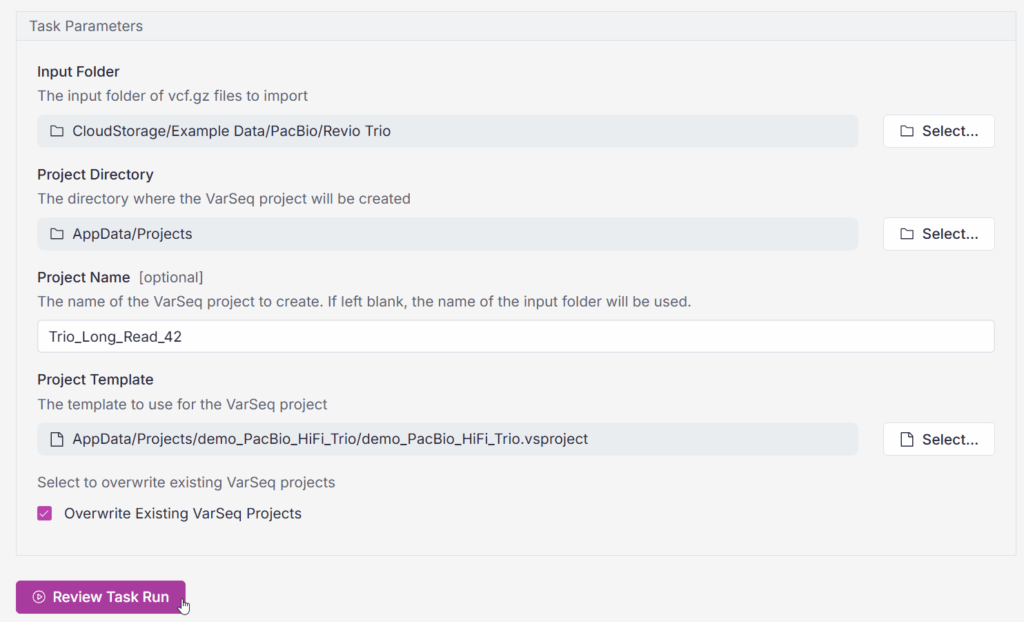
What exactly is a “task” in this context? Tasks are the highly configurable building blocks of bioinformatic workflows in VSWarehouse. The VSPipeline Create VarSeq Project task is illustrated in Figure 1. With one click on “Review Task Run,” this task is run on a cloud compute agent. The task automatically creates the batch script needed to run VSPipeline.
Automating Project Creation at Scale
Adding a VSPipeline Create VarSeq Project step to an existing pipeline is straightforward. By making project creation part of the workflow, scaling up does not add new bottlenecks for manual interpretation. Whether the lab is handling a handful of cases or many hundreds, the same VSPipeline automation can bridge secondary analysis (alignment and variant calling) with analysis of variants from gene panels, exomes, and whole genomes.
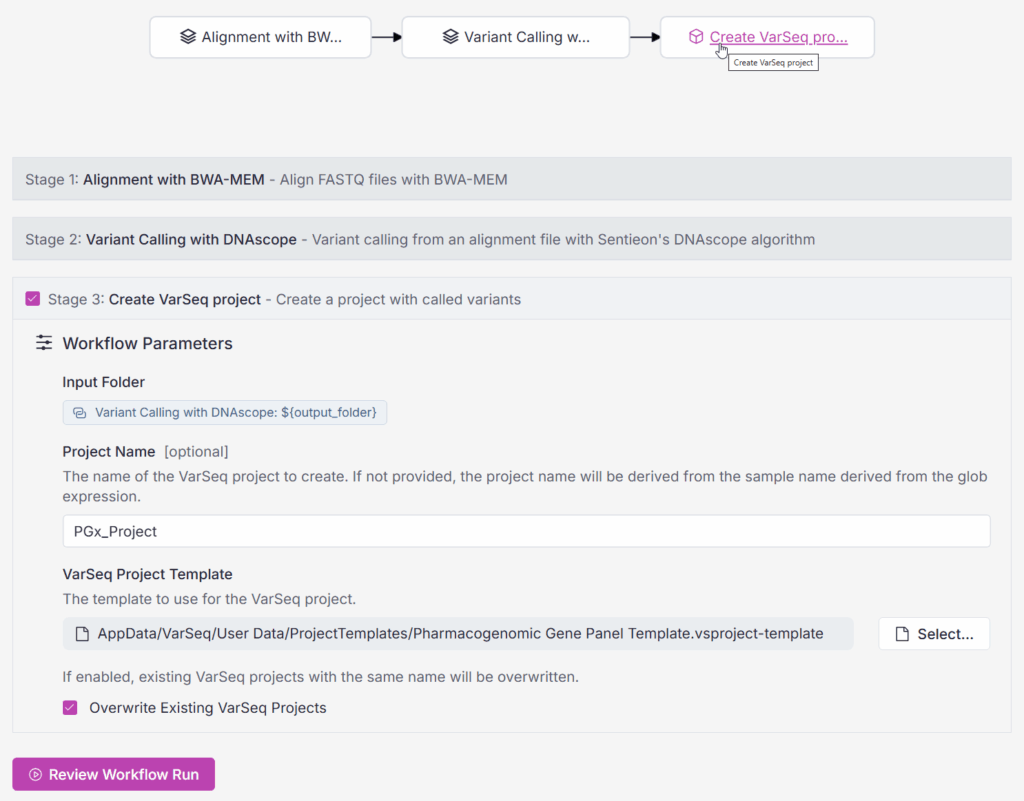
Figure 2 illustrates the incorporation of VSPipeline into a relatively simple workflow in VSWarehouse, which aligns and calls variants from short-read data using Sentieon before creating a VarSeq project. Note that the VSPipeline Create VarSeq Project task inherits the input folder information from previous steps in the workflow, so you don’t need to manually specify the inputs for each stage.
Comprehensive Use Case: PacBio WGS Variant Pipeline
For another more complex example, consider the PacBio WGS Variant Pipeline. We discussed implementing the singleton version of this workflow in a recent Webcast and the corresponding Blog Post. The PacBio WGS Singleton workflow consists of twelve modular stages, leveraging the parallel compute capability of VSWarehouse before funneling results into the final VSPipeline Create VarSeq Project task. This comprehensive workflow jumps from raw PacBio WGS data to a VarSeq project that includes SNVs, indels, tandem repeats, CNVs, and structural variants, in phased haplotypes. You can even view methylation patterns in the resulting VarSeq project via GenomeBrowse.
It’s important to remember that the benefits of automation in VSWarehouse do not detract from the potential for customization. If you want to customize VSPipeline tasks or upload your already-polished pipeline scripts to VSWarehouse, you can! Moreover, you can easily edit those scripts using VSCode within your VSWarehouse workspace, as shown in Figure 3.
VSPipeline is essential for guaranteeing the repeatability of clinical workflows, which is crucial from a regulatory point of view. It formalizes the steps between raw variant data and interpretation, so that every dataset is transformed the same way, with the right inputs, templates, and naming. Instead of spending time re-creating the same project structures, teams can rely on a predictable process that produces well-formed projects every time.
Automatically convert raw data into results with VSPipeline
VSPipeline makes the path from variant data to interpretation smoother and more predictable, and it’s more accessible than ever in VSWarehouse. By folding VarSeq project creation into the workflows that power your secondary analysis, VSPipeline removes repetitive setup steps and ensures every project is built from the same blueprint. For labs working at any scale, this ability to configure and automate tertiary analysis steps strikes a unique balance between adaptability and simplicity.
Want to learn more about how the Golden Helix software suite can streamline your workflows? Contact our team today!