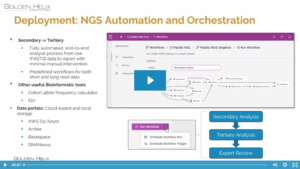
Using VSPipeline to Automate Tertiary Analysis in VSWarehouse

In this blog post, I’ll highlight the benefits of using VSPipeline to automate tertiary analysis in VSWarehouse. Users can now run VSPipeline project creation as a standalone task or a final step in a bioinformatic workflow without writing code. Simply use one of our shipped VarSeq projects or create your…
Read more →