Have you seen us in The Journal of Precision Medicine? Last week, our team released VarSeq 2.2.2 loaded with a number of updates and VSClinical’s highly-anticipated ACMG-CNV Guideline workflow! We have spent the past several months sharing webcasts and blogs on this new capability. We are honored to also have this new solution recognized in The Journal of Precision Medicine… Read more »
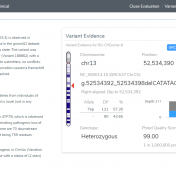
In our previous webcast, Evaluating CNVs with VSClinical’s New ACMG Guidelines, we focused on a CNV deletion (12:27715515-29628122×1) in which the patient had a known disorder called Brachydactyly type E. The CNV was isolated using our VS-CNV caller and applied to the ACMG CNV guidelines using the intuitive steps of VSClinical. If you missed the webcast, you can watch the… Read more »
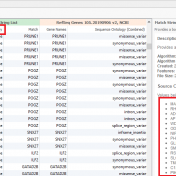
A common discussion with our customers includes the challenges with the tertiary stage of analyzing next-gen sequencing data. This is the stage where all data from gene panels, exome, or whole genome scale pass through filters to quickly isolate the clinically relevant variant contributing to a patient disorder. Golden Helix has recognized these challenges in the scale of data and… Read more »
It doesn’t take much effort to find articles discussing the value of Next-Generation Sequencing (NGS). There is a consistent tone amongst authors that implementing NGS pipelines are critical for clinical efficiency in both hereditary disorders and somatic. However, NGS strategies do not come without their own challenges. Challenges include not only the detection and calling of high quality/probability variants from… Read more »
It is an honor to be published in The Journal of Precision Medicine’s June/July 2020 issue in an article co-authored by Dr. Christiane Scherer and myself, “Diagnosing and Tracking COVID-19 Infections Leveraging Next-Gen Sequencing.” In this article, I detail: The key facts about COVID-19, reviewing the epidemiology, reservoir hosts, transmission routes, and clinical manifestation. Answering the question of how Next-Generation Sequencing can… Read more »
It is an honor to be published in The Journal of Precision Medicine’s March 2020 issue where I have outlined the current best practices for NGS-based cancer diagnostics. In this article, I detail: The need to standardize clinical reporting Popular annotation sources and functional prediction algorithms The AMP Guidelines and how these can be applied with Golden Helix’s Diagnostic Platform… Read more »
We have released a new version of my eBook, “Secondary Analysis 3.0”. To download a complimentary copy, I encourage you to do so here. Next-Generation Sequencing (NGS) promises to be the ultimate paradigm when it comes to genetic research and clinical testing since it contains the complete genetic information. When it comes to the current reality in testing labs, there… Read more »
We just got back from three busy days at the Molecular Pathology (AMP) conference in friendly San Antonio, Texas. Keeping up the Golden Helix conference momentum for the year, we had 3-4 in-booth demonstrations a day covering our CNV calling, variant interpretation, and data warehousing products for NGS-based genetic tests. And in short, NGS based tests for cancer and germline… Read more »
In 1914 the German cytologist Theodor Boveri coined the phrase “Cancer is a disease of the genome”. At this time his ideas were equally revolutionary as they were highly contested. Fast forward. More than hundred years later, Next-Generation Sequencing effectively permits a highly sensitive analysis of cancer cells. It can help us to understand mutations associated with cancer development and… Read more »