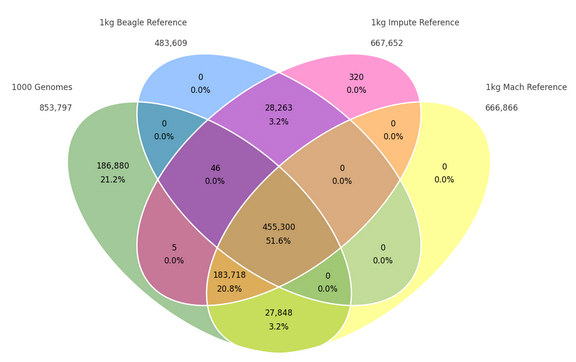
One frequent question I hear from SVS customers is whether whole exome sequence data can be used for principal components analysis (PCA) and other applications in population genetics. The answer is, “yes, but you need to be cautious.” What does cautious mean? Let’s take a look at the 1000 Genomes project for some examples.
There is a lot we can be grateful for at Golden Helix. The past year was marked by two major breakthrough launches. Earlier in 2014, we shipped SVS 8 which unified SVS with our GenomeBrowse product. We were able to improve SVS’ data management and visualization capabilities. In addition we added a number of new methods in SVS, such as SKAT-O, MM-KBAC, and various genomic prediction algorithms.
The Golden Helix team enjoys following our customers’ success. And we would like to share some recent client work to demonstrate what is possible with our software, as well as to inspire researchers to continue questioning current scientific norms.
We are shortly approaching the public launch (November 5th!) of our first clinical product, VarSeq. We could not have predicted how well the market would accept VarSeq – but we couldn’t be happier! For those of you who have not yet seen our newest product in action, I invite you to register for tomorrow’s webcast: The Golden Helix VarSeq User Experience.
Golden Helix is proud to announce the release of the Golden Helix GenomeBrowse Plugin for Ion Torrent server. The new plug-in enables adding selected BAM files from Torrent Server reports directly into GenomeBrowse. The BAM files remain on the torrent server and are streamed from the server on demand using your credentials. This feature allows GenomeBrowse users to visualize genomic… Read more »
Recently, I have been thinking a lot about Human Genome Variation Society (HGVS) notation — you know “G dot”, “P dot”, and “C dot”. HGVS has quickly become one of the most common ways to represent variants. It’s no wonder that HGVS nomenclature is used so widely. It provides an easily readable, compact representation of a variant. Since it is… Read more »
Last week, our CEO Andreas Scherer announced our entrance into the clinical testing market with VarSeq. This week, I will be giving a webcast on Wednesday introducing this new tool and demonstrating its capabilities. (Register for the webcast) VarSeq’s focused purpose is making NGS gene testing and variant discovery efficient, scalable and accessible to users with a broad range of backgrounds and specialties. In this blog post, we will examine the use cases that VarSeq supports in more detail,… Read more »
Over the last decade, DNA sequencing has made vast technological improvements. With the cost of sequencing decreasing significantly, sequencing technology has become a product for the masses. The sequencing technology and programs that were once used exclusively by major research institutions are now becoming available in many research facilities around the globe. These tools produce large amounts of data sets… Read more »
You probably haven’t spent much time thinking about how we represent genes in a genomic reference sequence context. And by genes, I really mean transcripts since genes are just a collection of transcripts that produce the same product. But in fact, there is more complexity here than you ever really wanted to know about. Andrew Jesaitis covered some of this… Read more »
We released GenomeBrowse 2.0 earlier this year, allowing users to review all types of genomic data. Since then, it has received rave reviews from thousands of users around the world. Essentially, it’s the Google Earth app for genomic data. GenomeBrowse allows a user to sift through vast amounts of genomic data, and make it easy to focus on a single part… Read more »
Up until a few weeks ago, I thought variant classification was basically a solved problem. I mean, how hard can it be? We look at variants all the time and say things like, “Well that one is probably not too detrimental since it’s a 3 base insertion, but this frameshift is worth looking into.” What we fail to recognize is… Read more »
When many people think of learning disabilities such as dyslexia and language impairment, they typically do not think of a biological or medical condition. Even more rarely do people think of these conditions as being the result of biological and genetic phenomena. However, that is exactly what I have thought of every day during my doctoral training in the Department… Read more »
Philadelphia. The City of Brotherly love, home of the Philadelphia cheesesteak, and home to the Children’s Hospital of Philadelphia, who took the No. 1 spot in this year’s U.S. News & World Report’s Best Children’s Hospitals Honor Roll. We are honored (and excited!) to sponsor this year’s Mid-Atlantic Epidemiology and Statistics (MAGES) Conference at the University of Pennsylvania. MAGES will… Read more »
As I write this article, Golden Helix has hundreds of clients in top research institutions world-wide. The adoption of our product at these institutions ranges from a few individual users to site licenses used by entire organizations. Because of the quality of SNP & Variation Suite (SVS) and GenomeBrowse, our competence in the field is recognized, and increasingly our clients… Read more »
On my flight back from this year’s Molecular Tri-Conference in San Francisco, I couldn’t help but ruminate over the intriguing talks, engaging round table discussions, and fabulous dinners with fellow speakers. And I kept returning to the topic of how we aggregate, share, and update data in the interest of understanding our genomes. Of course, there were many examples of… Read more »
A report from the World Congress of Psychiatric Genetics Earlier this month, while much of the genetics community was scrambling to edit and print their posters for ASHG, I had the opportunity to attend WCPG, the World Congress of Psychiatric Genetics, in Boston. This was my second trip to WCPG and it is becoming one of my favorite events to… Read more »
Genotype imputation is a common and useful practice that allows GWAS researchers to analyze untyped SNPs without the cost of genotyping millions of additional SNPs. In the Services Department at Golden Helix, we often perform imputation on client data, and we have our own software preferences for a variety of reasons. However, other imputation software packages have their own advantages… Read more »
In a couple of short weeks, Gabe is headed off to TCGC in San Francisco where he will be giving part of a short course. He was super excited about it last year and is even more so this year. I sat down with him yesterday to find out why. Jessica: What’s TCGC? Gabe: Last year I got to attend… Read more »
When researchers realized they needed a way to report genetic variants in scientific literature using a consistent format, the Human Genome Variation Society (HGVS) mutation nomenclature was developed and quickly became the standard method for describing sequence variations. Increasingly, HGVS nomenclature is being used to describe variants in genetic variant databases as well. There are some practical issues that researchers… Read more »
A few months ago, our CEO, Christophe Lambert, directed me toward an interesting commentary published in Nature Reviews Genetics by authors Bjarni J. Vilhjalmsson and Magnus Nordborg. Population structure is frequently cited as a major source of confounding in GWAS, but the authors of the article suggest that the problems often blamed on population structure actually result from the environment… Read more »