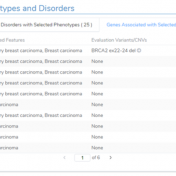
In this blog, we will be covering new assessment catalogs and how they work to improve saving and tracking variant interpretations. VarSeq is a variant analysis tool that effectively analyzes single nucleotide (SNVs) and copy number variants (CNVs) in both cancer and germline workflows. Because VarSeq enables such diverse variant analysis, there are many research labs and institutions that evaluate… Read more »
Orphanet is a public database available in VarSeq that aims to improve the current understanding and treatment options for patients affected by rare diseases. This resource was established in France in 1997 and has gradually grown to cover a consortium of over 40 countries in Europe. The primary goal of this database is to provide a universal nomenclature to classify… Read more »
We would like to thank everyone who entered our 2021 Abstract Competition. It is wonderful to read about the different ways Golden Helix software is being applied around the world. It is our pleasure to announce this year’s winners! First Place: Viney Gupta – All India Institute of Medical Sciences – AIIMS Digenic Inheritance in Juvenile Open-AngleGlaucoma Professor Viney Gupta… Read more »
Thank you to those who attended the recent webcast, “VSWarehouse: Tracking Changing Variant Evidence and Classifications”. For those who could not attend but wish to watch, here is a link to the recording. The webcast covered some general highlights of VSWarehouse value but also presented some specific capabilities covering the ClinVar classification tracker. Golden Helix provides complete solutions to handle… Read more »
Clinical testing labs produce reports as the end product of the NGS variant detection and interpretation workflow. Necessarily, the content, detail, and presentation of the report needs to be specialized to each clinical lab, and potentially each offered test. Our last blog post introduced the new Word-based report templates in VSClinical. In this blog post, we will introduce and explore… Read more »
The collaboration between the Clinical Genome Resource (ClinGen) consortium and the American College of Medical Genetics (ACMG) recently developed published guidelines for the interpretation of CNVs called on next-generation sequencing data. These new guidelines are the first to provide a robust set of rules for the interpretation of small intragenic deletions and duplications and are now automated in VSClinical. … Read more »
This morning I released a new version of my eBook “Clinical Variant Analysis – Second Edition.” The clinical interpretation of variants in Next-Gen Sequencing is a quickly evolving field. While the body of knowledge is growing exponentially, experts have to derive sound, clinical decisions leveraging an ever-expanding set of specialty databases, clinical publications, and algorithms that are designed to predict the… Read more »
In many cases, VarSeq users typically run single trio projects or perhaps an extended family project. Not only are all the inheritance model algorithms available in the VarSeq software to capture de novo, dominant, or recessively inherited variants but there are a number of quality control fields to help ensure the pedigree was set up properly. The last thing any… Read more »
Next-generation sequencing generates an immense amount of data which is then subject to a multi-step process to establish a validated bioinformatic pipeline. From processing raw sequence data to the detection of genetic mutations, establishing a validated and consistent bioinformatic pipeline makes a huge difference in the quality of patient care and accuracy of results. In this blog, we are focusing… Read more »
Welcome to our Customer Publications for February 2021, dedicated to cardiac research. As we commemorate the 57th American Heart Month, it is important to remember why we are wearing red and using #OurHearts. Heart disease is the leading cause of death worldwide. The National Heart, Lung, and Blood Institute urges Americans to do what they can to be heart-healthy and… Read more »
The recent release of VarSeq 2.2.2 brings our Word report template system, previously featured in VSClinical AMP, to the VSClinical ACMG workflow. This blog post will describe how to use the Word template system using one of our shipped templates as well as how to start customizing your own templates. We will cover the three different report templates that ship… Read more »
Our latest release of the VarSeq software has had a major upgrade with the addition of the new CNV ACMG guidelines! Here are some recent webcasts we’ve given covering the new guideline tool: Family-Based Workflows in VarSeq and VSClinical A User’s Perspective: ACMG Guidelines for CNVs in VSClinical Not only does VarSeq 2.2.2 come with the new guideline tool, but… Read more »
VarSeq recently received major upgrades in a wide range of areas, one of these areas includes adding annotations such as GnomAD. This includes new fundamental methods of CNV ACMG guideline processing but also a large number of small additions in annotations. One addition is the application of gnomADs – Gene Constraints. This provides various metrics for pathogenicity on a per… Read more »
VarSeq 2.2.2 has incorporated a multitude of interesting new features. In this blog, I want to continue discussing these features and how each can be incorporated into your workflow. I will also discuss the application of the Probability Segregation algorithm for copy number variation (CNV) analysis. The Probability Segregation algorithm is a new algorithm that has been added to VarSeq… Read more »
Our latest VarSeq release is one of the largest we’ve ever had, boasting an extensive list of new features and improvements. As part of this release, we have dramatically expanded our support for splice site analysis. This includes improvements to our novel splice site algorithm and support for splice site effect prediction along with several other small improvements. Novel Splice… Read more »
New discoveries using NGS data analysis are never-ending and are pushing precision medicine to the forefront. In the January 2021 customer publication blog, I am focusing on our VarSeq software as investigators harness its power to perform a variety of investigational study designs. From cancer to inherited rare disease research, VarSeq is the rising star in research and diagnostic tools…. Read more »
I am pleased to share with you the official release of my updated eBook, “Precision Medicine“. Almost 2,500 years ago, Hippocrates captured one of the key principles underlying precision medicine. In the 21st century, we take an understanding of the individual characteristics of a person to a new level. By leveraging information about an individual’s genome, we are able to… Read more »
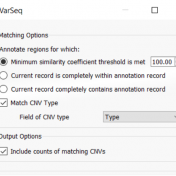
In continuation of our blog posts focusing on new features of VarSeq v2.2.2, here we will discuss the Latest Sample Assessment algorithm for both single nucleotide variants (SNVs) and copy number variants (CNVS). This algorithm annotates the variants of the project with the latest assessment from your variant catalog, which will show the history of interpretations made for the variants… Read more »
Annotating genomic variants is a very complex process but perhaps the most important part of next-generation sequencing variant analysis. Here at Golden Helix, we recognize the importance and value of having the most up-to-date sources available and curating new annotation sources as they become available for variant analysis. Golden Helix has curated over 100 annotation sources for human variant analysis… Read more »
Our recent release of VarSeq 2.2.2 comes with a long list of upgrades and new features. In this blog post, we will demonstrate how defining sample phenotypes are available in VSClinical. One noticeable change is the ACMG guideline variant evaluation in VSClinical. Not only has this interface added CNV guideline evaluation, simplified the reporting process with embedded Microsoft Word and… Read more »