Clinical Variant Analysis: Part V

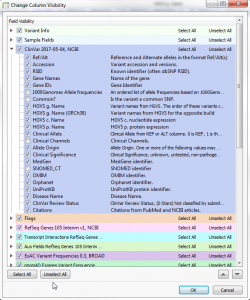
Examples of Clinical Variant Interpretation with VSClinical In this chapter, I’d like to go through a few examples for variants that have been classified with the help of VSClinical. This will give you a better understanding of how data sources are actually being represented in the software and how those…
Read more →