New Tutorial: VSReports

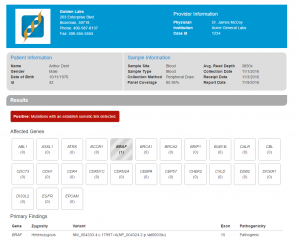
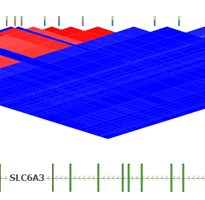
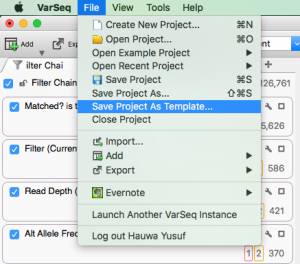
The new VSReports tutorial covers a basic VSReports workflow with an emphasis on understanding and exploring report customizations. This tutorial requires an active VarSeq license with the the VSReport feature included. You can go to Discover VarSeq or email [email protected] to request an evaluation license with the VSReports functionality included. VS Reports provides…
Read more →